1Goldberger, A., Amaral, L., Glass, L., Hausdorff, J., Ivanov, P. C., Mark, R., ... & Stanley, H. E. (2000). PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation Online. 101 (23), pp. e215--e220. RRID:SCR_007345.
SHA256SUMS.txt文件同样中记录了各个文件的命名,但需要调整细节,否则将报错:
Matlab
% 文件路径
filePath = 'D:\ECG-Tran\mit-bih-arrhythmia-database\SHA256SUMS.txt';
% 读取所有行
fid = fopen(filePath, 'r');
lines = textscan(fid, '%s', 'Delimiter', '\n', 'Whitespace', '');
fclose(fid);
lines = lines{1};
% 提取每行末尾的文件名
fileNames = {};
for i = 1:length(lines)
token = regexp(lines{i}, '\S+$', 'match', 'once');
if ~isempty(token)
fileNames{end+1} = token;
end
end
% 提取扩展名(包括短横线,如 '.hea-'),无扩展名的留空
extensions = cellfun(@(f) regexp(f, '\.[^.]*$', 'match', 'once'), fileNames, 'UniformOutput', false);
% 将空后缀替换为 'no_extension'
emptyIdx = cellfun(@isempty, extensions);
extensions(emptyIdx) = {'no_extension'};
% 获取唯一扩展名及其对应的合法字段名
uniqueExt = unique(extensions);
fieldNames = cellfun(@ext2fieldname, uniqueExt, 'UniformOutput', false);
% 分组并排序
sortedGroups = struct();
extOriginalMap = struct(); % 存储每个字段名对应的原始扩展名(用于显示)
for i = 1:length(uniqueExt)
ext = uniqueExt{i};
fieldName = fieldNames{i};
idx = strcmp(extensions, ext);
groupFiles = fileNames(idx);
% 排序:优先按文件名开头的数字升序,否则按字母序
nums = cellfun(@(f) regexp(f, '^\d+', 'match', 'once'), groupFiles, 'UniformOutput', false);
hasNum = ~cellfun(@isempty, nums);
if all(hasNum)
numVals = cellfun(@str2double, nums);
[~, sortIdx] = sort(numVals);
else
[~, sortIdx] = sort(groupFiles);
end
sortedGroups.(fieldName) = groupFiles(sortIdx);
extOriginalMap.(fieldName) = ext; % 保存原始扩展名
% 保存为 MAT 文件
save('sorted_filenames_by_ext.mat', 'sortedGroups', 'extOriginalMap');
end
% 显示结果(使用原始扩展名展示)
extFieldNames = fieldnames(sortedGroups);
for i = 1:length(extFieldNames)
fieldName = extFieldNames{i};
originalExt = extOriginalMap.(fieldName);
if strcmp(originalExt, 'no_extension')
displayExt = '无扩展名';
else
displayExt = originalExt;
end
fprintf('后缀: %s, 共 %d 个文件\n', displayExt, length(sortedGroups.(fieldName)));
disp(sortedGroups.(fieldName)');
end
% 将扩展名转换为合法的结构体字段名(例如 '.atr' -> 'atr', '.hea-' -> 'hea_')
function validName = ext2fieldname(ext)
if strcmp(ext, 'no_extension')
validName = 'no_extension';
return;
end
% 去掉开头的点号
if startsWith(ext, '.')
ext = ext(2:end);
end
% 将非字母数字下划线替换为下划线(例如 '-' -> '_')
validName = regexprep(ext, '[^a-zA-Z0-9_]', '_');
% 如果结果为空或首字符是数字,则添加前缀 'ext_'
if isempty(validName) || ismember(validName(1), '0123456789')
validName = ['ext_' validName];
end
end这里的文件相较于前文使用的数据库要更多了:
Matlab
后缀: .1, 共 1 个文件
{'mitdbdir/src/tab.1'}
后缀: .2, 共 1 个文件
{'mitdbdir/src/tab.2'}
后缀: .3, 共 1 个文件
{'mitdbdir/src/tab.3'}
后缀: .4, 共 1 个文件
{'mitdbdir/src/tab.4'}
后缀: .5, 共 1 个文件
{'mitdbdir/src/tab.5'}
后缀: .at_, 共 8 个文件
{'108.at_'}
{'117.at_'}
{'119.at_'}
{'203.at_'}
{'209.at_'}
{'214.at_'}
{'215.at_'}
{'222.at_'}
后缀: .atr, 共 72 个文件
{'100.atr' }
{'101.atr' }
{'102-0.atr' }
{'102.atr' }
{'103.atr' }
{'104.atr' }
{'105.atr' }
{'106.atr' }
{'107.atr' }
{'108.atr' }
{'109.atr' }
{'111.atr' }
{'112.atr' }
{'113.atr' }
{'114.atr' }
{'115.atr' }
{'116.atr' }
{'117.atr' }
{'118.atr' }
{'119.atr' }
{'121.atr' }
{'122.atr' }
{'123.atr' }
{'124.atr' }
{'200.atr' }
{'201.atr' }
{'202.atr' }
{'203.atr' }
{'205.atr' }
{'207.atr' }
{'208.atr' }
{'209.atr' }
{'210.atr' }
{'212.atr' }
{'213.atr' }
{'214.atr' }
{'215.atr' }
{'217.atr' }
{'219.atr' }
{'220.atr' }
{'221.atr' }
{'222.atr' }
{'223.atr' }
{'228.atr' }
{'230.atr' }
{'231.atr' }
{'232.atr' }
{'233.atr' }
{'234.atr' }
{'x_mitdb/x_108.atr'}
{'x_mitdb/x_109.atr'}
{'x_mitdb/x_111.atr'}
{'x_mitdb/x_112.atr'}
{'x_mitdb/x_113.atr'}
{'x_mitdb/x_114.atr'}
{'x_mitdb/x_115.atr'}
{'x_mitdb/x_116.atr'}
{'x_mitdb/x_117.atr'}
{'x_mitdb/x_121.atr'}
{'x_mitdb/x_122.atr'}
{'x_mitdb/x_123.atr'}
{'x_mitdb/x_124.atr'}
{'x_mitdb/x_220.atr'}
{'x_mitdb/x_221.atr'}
{'x_mitdb/x_222.atr'}
{'x_mitdb/x_223.atr'}
{'x_mitdb/x_228.atr'}
{'x_mitdb/x_230.atr'}
{'x_mitdb/x_231.atr'}
{'x_mitdb/x_232.atr'}
{'x_mitdb/x_233.atr'}
{'x_mitdb/x_234.atr'}
后缀: .bat, 共 1 个文件
{'mitdbdir/src/printdir.bat'}
后缀: .c, 共 3 个文件
{'mitdbdir/src/dbnotes-html.c'}
{'mitdbdir/src/dbnotes.c' }
{'mitdbdir/src/dbtab.c' }
后缀: .dat, 共 71 个文件
{'100.dat' }
{'101.dat' }
{'102.dat' }
{'103.dat' }
{'104.dat' }
{'105.dat' }
{'106.dat' }
{'107.dat' }
{'108.dat' }
{'109.dat' }
{'111.dat' }
{'112.dat' }
{'113.dat' }
{'114.dat' }
{'115.dat' }
{'116.dat' }
{'117.dat' }
{'118.dat' }
{'119.dat' }
{'121.dat' }
{'122.dat' }
{'123.dat' }
{'124.dat' }
{'200.dat' }
{'201.dat' }
{'202.dat' }
{'203.dat' }
{'205.dat' }
{'207.dat' }
{'208.dat' }
{'209.dat' }
{'210.dat' }
{'212.dat' }
{'213.dat' }
{'214.dat' }
{'215.dat' }
{'217.dat' }
{'219.dat' }
{'220.dat' }
{'221.dat' }
{'222.dat' }
{'223.dat' }
{'228.dat' }
{'230.dat' }
{'231.dat' }
{'232.dat' }
{'233.dat' }
{'234.dat' }
{'x_mitdb/x_108.dat'}
{'x_mitdb/x_109.dat'}
{'x_mitdb/x_111.dat'}
{'x_mitdb/x_112.dat'}
{'x_mitdb/x_113.dat'}
{'x_mitdb/x_114.dat'}
{'x_mitdb/x_115.dat'}
{'x_mitdb/x_116.dat'}
{'x_mitdb/x_117.dat'}
{'x_mitdb/x_121.dat'}
{'x_mitdb/x_122.dat'}
{'x_mitdb/x_123.dat'}
{'x_mitdb/x_124.dat'}
{'x_mitdb/x_220.dat'}
{'x_mitdb/x_221.dat'}
{'x_mitdb/x_222.dat'}
{'x_mitdb/x_223.dat'}
{'x_mitdb/x_228.dat'}
{'x_mitdb/x_230.dat'}
{'x_mitdb/x_231.dat'}
{'x_mitdb/x_232.dat'}
{'x_mitdb/x_233.dat'}
{'x_mitdb/x_234.dat'}
后缀: .dvi, 共 1 个文件
{'mitdbdir/src/title.dvi'}
后缀: .gz, 共 9 个文件
{'mitdbdir/src/contents.gz'}
{'mitdbdir/src/cover.gz' }
{'mitdbdir/src/extext.gz' }
{'mitdbdir/src/fdtext.gz' }
{'mitdbdir/src/index.gz' }
{'mitdbdir/src/intro.gz' }
{'mitdbdir/src/notes.gz' }
{'mitdbdir/src/tables.gz' }
{'mitdbdir/src/title.gz' }
后缀: .h, 共 1 个文件
{'mitdbdir/src/notes.h'}
后缀: .hea, 共 71 个文件
{'100.hea' }
{'101.hea' }
{'102.hea' }
{'103.hea' }
{'104.hea' }
{'105.hea' }
{'106.hea' }
{'107.hea' }
{'108.hea' }
{'109.hea' }
{'111.hea' }
{'112.hea' }
{'113.hea' }
{'114.hea' }
{'115.hea' }
{'116.hea' }
{'117.hea' }
{'118.hea' }
{'119.hea' }
{'121.hea' }
{'122.hea' }
{'123.hea' }
{'124.hea' }
{'200.hea' }
{'201.hea' }
{'202.hea' }
{'203.hea' }
{'205.hea' }
{'207.hea' }
{'208.hea' }
{'209.hea' }
{'210.hea' }
{'212.hea' }
{'213.hea' }
{'214.hea' }
{'215.hea' }
{'217.hea' }
{'219.hea' }
{'220.hea' }
{'221.hea' }
{'222.hea' }
{'223.hea' }
{'228.hea' }
{'230.hea' }
{'231.hea' }
{'232.hea' }
{'233.hea' }
{'234.hea' }
{'x_mitdb/x_108.hea'}
{'x_mitdb/x_109.hea'}
{'x_mitdb/x_111.hea'}
{'x_mitdb/x_112.hea'}
{'x_mitdb/x_113.hea'}
{'x_mitdb/x_114.hea'}
{'x_mitdb/x_115.hea'}
{'x_mitdb/x_116.hea'}
{'x_mitdb/x_117.hea'}
{'x_mitdb/x_121.hea'}
{'x_mitdb/x_122.hea'}
{'x_mitdb/x_123.hea'}
{'x_mitdb/x_124.hea'}
{'x_mitdb/x_220.hea'}
{'x_mitdb/x_221.hea'}
{'x_mitdb/x_222.hea'}
{'x_mitdb/x_223.hea'}
{'x_mitdb/x_228.hea'}
{'x_mitdb/x_230.hea'}
{'x_mitdb/x_231.hea'}
{'x_mitdb/x_232.hea'}
{'x_mitdb/x_233.hea'}
{'x_mitdb/x_234.hea'}
后缀: .htm, 共 5 个文件
{'mitdbdir/foreword.htm'}
{'mitdbdir/intro.htm' }
{'mitdbdir/mitdbdir.htm'}
{'mitdbdir/records.htm' }
{'mitdbdir/tables.htm' }
后缀: .pl, 共 1 个文件
{'mitdbdir/src/makelinks.pl'}
后缀: .tr, 共 6 个文件
{'mitdbdir/src/contents.tr'}
{'mitdbdir/src/cover.tr' }
{'mitdbdir/src/extext.tr' }
{'mitdbdir/src/fdtext.tr' }
{'mitdbdir/src/index.tr' }
{'mitdbdir/src/intro.tr' }
后缀: .xws, 共 439 个文件
{'100.xws' }
{'101.xws' }
{'102.xws' }
{'103.xws' }
{'104.xws' }
{'105.xws' }
{'106.xws' }
{'107.xws' }
{'108.xws' }
{'109.xws' }
{'111.xws' }
{'112.xws' }
{'113.xws' }
{'114.xws' }
{'115.xws' }
{'116.xws' }
{'117.xws' }
{'118.xws' }
{'119.xws' }
{'121.xws' }
{'122.xws' }
{'123.xws' }
{'124.xws' }
{'200.xws' }
{'201.xws' }
{'202.xws' }
{'203.xws' }
{'205.xws' }
{'207.xws' }
{'208.xws' }
{'209.xws' }
{'210.xws' }
{'212.xws' }
{'213.xws' }
{'214.xws' }
{'215.xws' }
{'217.xws' }
{'219.xws' }
{'220.xws' }
{'221.xws' }
{'222.xws' }
{'223.xws' }
{'228.xws' }
{'230.xws' }
{'231.xws' }
{'232.xws' }
{'233.xws' }
{'234.xws' }
{'mitdbdir/samples/1001103.xws'}
{'mitdbdir/samples/1002513.xws'}
{'mitdbdir/samples/1002609.xws'}
{'mitdbdir/samples/1002755.xws'}
{'mitdbdir/samples/1010134.xws'}
{'mitdbdir/samples/1010148.xws'}
{'mitdbdir/samples/1010513.xws'}
{'mitdbdir/samples/1010954.xws'}
{'mitdbdir/samples/1012432.xws'}
{'mitdbdir/samples/1020055.xws'}
{'mitdbdir/samples/1020112.xws'}
{'mitdbdir/samples/1020128.xws'}
{'mitdbdir/samples/1020230.xws'}
{'mitdbdir/samples/1020451.xws'}
{'mitdbdir/samples/1020935.xws'}
{'mitdbdir/samples/1021612.xws'}
{'mitdbdir/samples/1030109.xws'}
{'mitdbdir/samples/1031721.xws'}
{'mitdbdir/samples/1031915.xws'}
{'mitdbdir/samples/1032213.xws'}
{'mitdbdir/samples/1032333.xws'}
{'mitdbdir/samples/1032858.xws'}
{'mitdbdir/samples/1040342.xws'}
{'mitdbdir/samples/1040513.xws'}
{'mitdbdir/samples/1040552.xws'}
{'mitdbdir/samples/1040617.xws'}
{'mitdbdir/samples/1040822.xws'}
{'mitdbdir/samples/1042651.xws'}
{'mitdbdir/samples/1042910.xws'}
{'mitdbdir/samples/1050527.xws'}
{'mitdbdir/samples/1050757.xws'}
{'mitdbdir/samples/1051516.xws'}
{'mitdbdir/samples/1051752.xws'}
{'mitdbdir/samples/1052202.xws'}
{'mitdbdir/samples/1052645.xws'}
{'mitdbdir/samples/1052727.xws'}
{'mitdbdir/samples/1052808.xws'}
{'mitdbdir/samples/1052907.xws'}
{'mitdbdir/samples/1060019.xws'}
{'mitdbdir/samples/1060137.xws'}
{'mitdbdir/samples/1060253.xws'}
{'mitdbdir/samples/1060423.xws'}
{'mitdbdir/samples/1060757.xws'}
{'mitdbdir/samples/1061052.xws'}
{'mitdbdir/samples/1061227.xws'}
{'mitdbdir/samples/1061617.xws'}
{'mitdbdir/samples/1062513.xws'}
{'mitdbdir/samples/1062552.xws'}
{'mitdbdir/samples/1070044.xws'}
{'mitdbdir/samples/1071230.xws'}
{'mitdbdir/samples/1071954.xws'}
{'mitdbdir/samples/1072038.xws'}
{'mitdbdir/samples/1072552.xws'}
{'mitdbdir/samples/1080022.xws'}
{'mitdbdir/samples/1080451.xws'}
{'mitdbdir/samples/1080741.xws'}
{'mitdbdir/samples/1080813.xws'}
{'mitdbdir/samples/1081055.xws'}
{'mitdbdir/samples/1081808.xws'}
{'mitdbdir/samples/1082005.xws'}
{'mitdbdir/samples/1082420.xws'}
{'mitdbdir/samples/1082810.xws'}
{'mitdbdir/samples/1082900.xws'}
{'mitdbdir/samples/1090013.xws'}
{'mitdbdir/samples/1090128.xws'}
{'mitdbdir/samples/1090336.xws'}
{'mitdbdir/samples/1090446.xws'}
{'mitdbdir/samples/1090527.xws'}
{'mitdbdir/samples/1091401.xws'}
{'mitdbdir/samples/1091713.xws'}
{'mitdbdir/samples/1091921.xws'}
{'mitdbdir/samples/1092609.xws'}
{'mitdbdir/samples/1092803.xws'}
{'mitdbdir/samples/1092828.xws'}
{'mitdbdir/samples/1092910.xws'}
{'mitdbdir/samples/1110025.xws'}
{'mitdbdir/samples/1110241.xws'}
{'mitdbdir/samples/1110352.xws'}
{'mitdbdir/samples/1110404.xws'}
{'mitdbdir/samples/1110831.xws'}
{'mitdbdir/samples/1111153.xws'}
{'mitdbdir/samples/1111541.xws'}
{'mitdbdir/samples/1111954.xws'}
{'mitdbdir/samples/1112702.xws'}
{'mitdbdir/samples/1121140.xws'}
{'mitdbdir/samples/1121230.xws'}
{'mitdbdir/samples/1121249.xws'}
{'mitdbdir/samples/1130420.xws'}
{'mitdbdir/samples/1130822.xws'}
{'mitdbdir/samples/1131148.xws'}
{'mitdbdir/samples/1131227.xws'}
{'mitdbdir/samples/1132210.xws'}
{'mitdbdir/samples/1132901.xws'}
{'mitdbdir/samples/1140000.xws'}
{'mitdbdir/samples/1140120.xws'}
{'mitdbdir/samples/1140339.xws'}
{'mitdbdir/samples/1140356.xws'}
{'mitdbdir/samples/1140435.xws'}
{'mitdbdir/samples/1140536.xws'}
{'mitdbdir/samples/1140831.xws'}
{'mitdbdir/samples/1140926.xws'}
{'mitdbdir/samples/1141137.xws'}
{'mitdbdir/samples/1142002.xws'}
{'mitdbdir/samples/1142929.xws'}
{'mitdbdir/samples/1150055.xws'}
{'mitdbdir/samples/1150322.xws'}
{'mitdbdir/samples/1151552.xws'}
{'mitdbdir/samples/1152128.xws'}
{'mitdbdir/samples/1152705.xws'}
{'mitdbdir/samples/1152842.xws'}
{'mitdbdir/samples/1160044.xws'}
{'mitdbdir/samples/1160131.xws'}
{'mitdbdir/samples/1161000.xws'}
{'mitdbdir/samples/1161232.xws'}
{'mitdbdir/samples/1161637.xws'}
{'mitdbdir/samples/1162308.xws'}
{'mitdbdir/samples/1162547.xws'}
{'mitdbdir/samples/1170356.xws'}
{'mitdbdir/samples/1171158.xws'}
{'mitdbdir/samples/1172227.xws'}
{'mitdbdir/samples/1180339.xws'}
{'mitdbdir/samples/1180831.xws'}
{'mitdbdir/samples/1180923.xws'}
{'mitdbdir/samples/1181347.xws'}
{'mitdbdir/samples/1182232.xws'}
{'mitdbdir/samples/1182325.xws'}
{'mitdbdir/samples/1182541.xws'}
{'mitdbdir/samples/1182623.xws'}
{'mitdbdir/samples/1182648.xws'}
{'mitdbdir/samples/1182858.xws'}
{'mitdbdir/samples/1182953.xws'}
{'mitdbdir/samples/1190155.xws'}
{'mitdbdir/samples/1190238.xws'}
{'mitdbdir/samples/1190451.xws'}
{'mitdbdir/samples/1190842.xws'}
{'mitdbdir/samples/1192005.xws'}
{'mitdbdir/samples/1192533.xws'}
{'mitdbdir/samples/1210106.xws'}
{'mitdbdir/samples/1210333.xws'}
{'mitdbdir/samples/1211648.xws'}
{'mitdbdir/samples/1212432.xws'}
{'mitdbdir/samples/1212601.xws'}
{'mitdbdir/samples/1220109.xws'}
{'mitdbdir/samples/1221347.xws'}
{'mitdbdir/samples/1222825.xws'}
{'mitdbdir/samples/1232210.xws'}
{'mitdbdir/samples/1232511.xws'}
{'mitdbdir/samples/1232741.xws'}
{'mitdbdir/samples/1232836.xws'}
{'mitdbdir/samples/1240443.xws'}
{'mitdbdir/samples/1240509.xws'}
{'mitdbdir/samples/1240521.xws'}
{'mitdbdir/samples/1240905.xws'}
{'mitdbdir/samples/1241027.xws'}
{'mitdbdir/samples/1241738.xws'}
{'mitdbdir/samples/1242013.xws'}
{'mitdbdir/samples/1242322.xws'}
{'mitdbdir/samples/1242603.xws'}
{'mitdbdir/samples/1242741.xws'}
{'mitdbdir/samples/2000142.xws'}
{'mitdbdir/samples/2000538.xws'}
{'mitdbdir/samples/2001814.xws'}
{'mitdbdir/samples/2002052.xws'}
{'mitdbdir/samples/2002449.xws'}
{'mitdbdir/samples/2002612.xws'}
{'mitdbdir/samples/2002831.xws'}
{'mitdbdir/samples/2002901.xws'}
{'mitdbdir/samples/2002918.xws'}
{'mitdbdir/samples/2002951.xws'}
{'mitdbdir/samples/2010000.xws'}
{'mitdbdir/samples/2010615.xws'}
{'mitdbdir/samples/2010752.xws'}
{'mitdbdir/samples/2010853.xws'}
{'mitdbdir/samples/2011814.xws'}
{'mitdbdir/samples/2012016.xws'}
{'mitdbdir/samples/2012233.xws'}
{'mitdbdir/samples/2012350.xws'}
{'mitdbdir/samples/2012415.xws'}
{'mitdbdir/samples/2012519.xws'}
{'mitdbdir/samples/2012620.xws'}
{'mitdbdir/samples/2012803.xws'}
{'mitdbdir/samples/2021016.xws'}
{'mitdbdir/samples/2021224.xws'}
{'mitdbdir/samples/2021241.xws'}
{'mitdbdir/samples/2021822.xws'}
{'mitdbdir/samples/2021845.xws'}
{'mitdbdir/samples/2021859.xws'}
{'mitdbdir/samples/2022110.xws'}
{'mitdbdir/samples/2022126.xws'}
{'mitdbdir/samples/2022213.xws'}
{'mitdbdir/samples/2022558.xws'}
{'mitdbdir/samples/2022755.xws'}
{'mitdbdir/samples/2022935.xws'}
{'mitdbdir/samples/2030500.xws'}
{'mitdbdir/samples/2031314.xws'}
{'mitdbdir/samples/2031502.xws'}
{'mitdbdir/samples/2032202.xws'}
{'mitdbdir/samples/2032325.xws'}
{'mitdbdir/samples/2032404.xws'}
{'mitdbdir/samples/2032446.xws'}
{'mitdbdir/samples/2032639.xws'}
{'mitdbdir/samples/2032651.xws'}
{'mitdbdir/samples/2032715.xws'}
{'mitdbdir/samples/2050457.xws'}
{'mitdbdir/samples/2051454.xws'}
{'mitdbdir/samples/2051522.xws'}
{'mitdbdir/samples/2051603.xws'}
{'mitdbdir/samples/2051615.xws'}
{'mitdbdir/samples/2051957.xws'}
{'mitdbdir/samples/2052418.xws'}
{'mitdbdir/samples/2052430.xws'}
{'mitdbdir/samples/2052757.xws'}
{'mitdbdir/samples/2070000.xws'}
{'mitdbdir/samples/2070033.xws'}
{'mitdbdir/samples/2070045.xws'}
{'mitdbdir/samples/2070057.xws'}
{'mitdbdir/samples/2070128.xws'}
{'mitdbdir/samples/2070207.xws'}
{'mitdbdir/samples/2070241.xws'}
{'mitdbdir/samples/2070425.xws'}
{'mitdbdir/samples/2070519.xws'}
{'mitdbdir/samples/2070656.xws'}
{'mitdbdir/samples/2071735.xws'}
{'mitdbdir/samples/2072536.xws'}
{'mitdbdir/samples/2072610.xws'}
{'mitdbdir/samples/2072714.xws'}
{'mitdbdir/samples/2072910.xws'}
{'mitdbdir/samples/2081457.xws'}
{'mitdbdir/samples/2081702.xws'}
{'mitdbdir/samples/2081910.xws'}
{'mitdbdir/samples/2082106.xws'}
{'mitdbdir/samples/2082300.xws'}
{'mitdbdir/samples/2082808.xws'}
{'mitdbdir/samples/2082858.xws'}
{'mitdbdir/samples/2082937.xws'}
{'mitdbdir/samples/2090047.xws'}
{'mitdbdir/samples/2090550.xws'}
{'mitdbdir/samples/2091257.xws'}
{'mitdbdir/samples/2091421.xws'}
{'mitdbdir/samples/2092325.xws'}
{'mitdbdir/samples/2092817.xws'}
{'mitdbdir/samples/2092833.xws'}
{'mitdbdir/samples/2092910.xws'}
{'mitdbdir/samples/2100347.xws'}
{'mitdbdir/samples/2100656.xws'}
{'mitdbdir/samples/2101314.xws'}
{'mitdbdir/samples/2101555.xws'}
{'mitdbdir/samples/2101732.xws'}
{'mitdbdir/samples/2101757.xws'}
{'mitdbdir/samples/2102033.xws'}
{'mitdbdir/samples/2102915.xws'}
{'mitdbdir/samples/2120005.xws'}
{'mitdbdir/samples/2121842.xws'}
{'mitdbdir/samples/2122421.xws'}
{'mitdbdir/samples/2122438.xws'}
{'mitdbdir/samples/2122645.xws'}
{'mitdbdir/samples/2122846.xws'}
{'mitdbdir/samples/2130339.xws'}
{'mitdbdir/samples/2131448.xws'}
{'mitdbdir/samples/2131505.xws'}
{'mitdbdir/samples/2131538.xws'}
{'mitdbdir/samples/2131730.xws'}
{'mitdbdir/samples/2131755.xws'}
{'mitdbdir/samples/2132443.xws'}
{'mitdbdir/samples/2132558.xws'}
{'mitdbdir/samples/2132651.xws'}
{'mitdbdir/samples/2132856.xws'}
{'mitdbdir/samples/2140030.xws'}
{'mitdbdir/samples/2140221.xws'}
{'mitdbdir/samples/2140235.xws'}
{'mitdbdir/samples/2140325.xws'}
{'mitdbdir/samples/2140356.xws'}
{'mitdbdir/samples/2140508.xws'}
{'mitdbdir/samples/2140538.xws'}
{'mitdbdir/samples/2141353.xws'}
{'mitdbdir/samples/2142317.xws'}
{'mitdbdir/samples/2142752.xws'}
{'mitdbdir/samples/2150255.xws'}
{'mitdbdir/samples/2150311.xws'}
{'mitdbdir/samples/2150946.xws'}
{'mitdbdir/samples/2151558.xws'}
{'mitdbdir/samples/2152025.xws'}
{'mitdbdir/samples/2152216.xws'}
{'mitdbdir/samples/2152457.xws'}
{'mitdbdir/samples/2152730.xws'}
{'mitdbdir/samples/2152757.xws'}
{'mitdbdir/samples/2152942.xws'}
{'mitdbdir/samples/2170000.xws'}
{'mitdbdir/samples/2170033.xws'}
{'mitdbdir/samples/2170123.xws'}
{'mitdbdir/samples/2170418.xws'}
{'mitdbdir/samples/2170612.xws'}
{'mitdbdir/samples/2170651.xws'}
{'mitdbdir/samples/2170705.xws'}
{'mitdbdir/samples/2171227.xws'}
{'mitdbdir/samples/2171401.xws'}
{'mitdbdir/samples/2171530.xws'}
{'mitdbdir/samples/2171656.xws'}
{'mitdbdir/samples/2172227.xws'}
{'mitdbdir/samples/2190249.xws'}
{'mitdbdir/samples/2190508.xws'}
{'mitdbdir/samples/2190522.xws'}
{'mitdbdir/samples/2191350.xws'}
{'mitdbdir/samples/2191900.xws'}
{'mitdbdir/samples/2191957.xws'}
{'mitdbdir/samples/2192211.xws'}
{'mitdbdir/samples/2192243.xws'}
{'mitdbdir/samples/2192443.xws'}
{'mitdbdir/samples/2192609.xws'}
{'mitdbdir/samples/2192855.xws'}
{'mitdbdir/samples/2200646.xws'}
{'mitdbdir/samples/2201041.xws'}
{'mitdbdir/samples/2201810.xws'}
{'mitdbdir/samples/2202029.xws'}
{'mitdbdir/samples/2202358.xws'}
{'mitdbdir/samples/2202544.xws'}
{'mitdbdir/samples/2202940.xws'}
{'mitdbdir/samples/2210000.xws'}
{'mitdbdir/samples/2210243.xws'}
{'mitdbdir/samples/2211300.xws'}
{'mitdbdir/samples/2211356.xws'}
{'mitdbdir/samples/2211749.xws'}
{'mitdbdir/samples/2211912.xws'}
{'mitdbdir/samples/2211942.xws'}
{'mitdbdir/samples/2212044.xws'}
{'mitdbdir/samples/2220645.xws'}
{'mitdbdir/samples/2221732.xws'}
{'mitdbdir/samples/2221948.xws'}
{'mitdbdir/samples/2222007.xws'}
{'mitdbdir/samples/2222232.xws'}
{'mitdbdir/samples/2222443.xws'}
{'mitdbdir/samples/2222543.xws'}
{'mitdbdir/samples/2222609.xws'}
{'mitdbdir/samples/2230935.xws'}
{'mitdbdir/samples/2231235.xws'}
{'mitdbdir/samples/2231339.xws'}
{'mitdbdir/samples/2231351.xws'}
{'mitdbdir/samples/2231721.xws'}
{'mitdbdir/samples/2231755.xws'}
{'mitdbdir/samples/2231907.xws'}
{'mitdbdir/samples/2232358.xws'}
{'mitdbdir/samples/2232547.xws'}
{'mitdbdir/samples/2232916.xws'}
{'mitdbdir/samples/2232951.xws'}
{'mitdbdir/samples/2280019.xws'}
{'mitdbdir/samples/2280038.xws'}
{'mitdbdir/samples/2280050.xws'}
{'mitdbdir/samples/2280435.xws'}
{'mitdbdir/samples/2281918.xws'}
{'mitdbdir/samples/2282008.xws'}
{'mitdbdir/samples/2282137.xws'}
{'mitdbdir/samples/2282149.xws'}
{'mitdbdir/samples/2282642.xws'}
{'mitdbdir/samples/2300000.xws'}
{'mitdbdir/samples/2300012.xws'}
{'mitdbdir/samples/2300511.xws'}
{'mitdbdir/samples/2300926.xws'}
{'mitdbdir/samples/2302904.xws'}
{'mitdbdir/samples/2310002.xws'}
{'mitdbdir/samples/2310224.xws'}
{'mitdbdir/samples/2312011.xws'}
{'mitdbdir/samples/2312110.xws'}
{'mitdbdir/samples/2312322.xws'}
{'mitdbdir/samples/2320058.xws'}
{'mitdbdir/samples/2320151.xws'}
{'mitdbdir/samples/2321151.xws'}
{'mitdbdir/samples/2321306.xws'}
{'mitdbdir/samples/2321509.xws'}
{'mitdbdir/samples/2321817.xws'}
{'mitdbdir/samples/2322231.xws'}
{'mitdbdir/samples/2322247.xws'}
{'mitdbdir/samples/2322320.xws'}
{'mitdbdir/samples/2322334.xws'}
{'mitdbdir/samples/2322353.xws'}
{'mitdbdir/samples/2330011.xws'}
{'mitdbdir/samples/2330025.xws'}
{'mitdbdir/samples/2330218.xws'}
{'mitdbdir/samples/2330243.xws'}
{'mitdbdir/samples/2330339.xws'}
{'mitdbdir/samples/2330502.xws'}
{'mitdbdir/samples/2330943.xws'}
{'mitdbdir/samples/2331620.xws'}
{'mitdbdir/samples/2331802.xws'}
{'mitdbdir/samples/2332223.xws'}
{'mitdbdir/samples/2340016.xws'}
{'mitdbdir/samples/2340637.xws'}
{'mitdbdir/samples/2341401.xws'}
{'mitdbdir/samples/2341426.xws'}
{'mitdbdir/samples/2341702.xws'}
{'mitdbdir/samples/2342126.xws'}
{'mitdbdir/samples/2342317.xws'}
后缀: 无扩展名, 共 10 个文件
{'ANNOTATORS' }
{'RECORDS' }
{'mitdbdir/src/domit' }
{'mitdbdir/src/exlist' }
{'mitdbdir/src/extest' }
{'mitdbdir/src/fdlist' }
{'mitdbdir/src/makefile' }
{'mitdbdir/src/makefile-html'}
{'x_mitdb/ANNOTATORS' }
{'x_mitdb/RECORDS' }但我发现一个问题:官方称含48 个充分标注的30分钟长度2导联ECG,但我们可以看出其有71个data文件。多了23个,具体是前缀多了个x,如:x_234.dat和234.dat,但没有资料说明有什么具体区别。
通过可视化观察一下:
Matlab
% 脚本:compare_x234_234.m
% 比较 x_234 和 234 记录经过 readdata 处理后的心电图信号
clear; close all;
% 数据文件夹路径
dataFolder = 'D:\ECG-Tran\mit-bih-arrhythmia-database\';
% 确保文件夹以文件分隔符结尾
if ~endsWith(dataFolder, filesep)
dataFolder = [dataFolder, filesep];
end
recordA = '234';
recordB = 'x_234';
fprintf('正在读取记录 %s ...\n', recordA);
try
[sigA, FsA] = readdata(recordA, dataFolder);
catch ME_A
warning('readdata 读取 %s 失败,改为直接读取原始信号。错误信息:%s', recordA, ME_A.message);
% 回退:读取原始完整信号
oldFolder = cd(dataFolder);
[sigA_raw, FsA, ~] = rdsamp(recordA);
cd(oldFolder);
sigA = sigA_raw; % 不进行任何预处理
end
fprintf('正在读取记录 %s ...\n', recordB);
try
[sigB, FsB] = readdata(recordB, dataFolder);
catch ME_B
warning('readdata 读取 %s 失败,改为直接读取原始信号。错误信息:%s', recordB, ME_B.message);
oldFolder = cd(dataFolder);
[sigB_raw, FsB, ~] = rdsamp(recordB);
cd(oldFolder);
sigB = sigB_raw;
end
% 检查采样率是否一致
if FsA ~= FsB
error('两个记录的采样率不一致,无法直接比较。');
end
Fs = FsA;
% 让比较窗口对齐:取两者中较短的信号长度
minLen = min(size(sigA,1), size(sigB,1));
sigA = sigA(1:minLen, :);
sigB = sigB(1:minLen, :);
% 生成统一的时间轴(单位:秒)
t = (0:minLen-1) / Fs;
% 只显示前 10 秒的数据,便于观察细节
viewSec = 10;
viewSamples = min(round(viewSec * Fs), minLen);
tv = t(1:viewSamples);
% ---- 可视化 ----
figure('Name', '信号对比:234 vs x_234', 'Position', [100, 100, 1200, 800]);
% 1. 第一导联叠加对比
subplot(3,1,1);
plot(tv, sigA(1:viewSamples, 1), 'b-', 'LineWidth', 1.2); hold on;
plot(tv, sigB(1:viewSamples, 1), 'r--', 'LineWidth', 1.2);
xlabel('时间 (秒)'); ylabel('幅值 (mV)');
title(sprintf('导联 1 前 %d 秒(蓝: 234, 红: x_234)', viewSec));
legend('234', 'x_234');
grid on;
% 2. 第一导联差异信号
diff_lead1 = sigA(1:viewSamples, 1) - sigB(1:viewSamples, 1);
subplot(3,1,2);
plot(tv, diff_lead1, 'k-', 'LineWidth', 1);
xlabel('时间 (秒)'); ylabel('差异 (mV)');
title('导联 1 差异 (234 - x_234)');
grid on;
% 添加零线
hold on; plot([tv(1), tv(end)], [0 0], 'Color', [0.6 0.6 0.6]);
% 3. 第二导联叠加对比
subplot(3,1,3);
plot(tv, sigA(1:viewSamples, 2), 'b-', 'LineWidth', 1.2); hold on;
plot(tv, sigB(1:viewSamples, 2), 'r--', 'LineWidth', 1.2);
xlabel('时间 (秒)'); ylabel('幅值 (mV)');
title(sprintf('导联 2 前 %d 秒(蓝: 234, 红: x_234)', viewSec));
legend('234', 'x_234');
grid on;
sgtitle(sprintf('记录 %s 与 %s 经过 readdata 处理后的信号对比', recordA, recordB));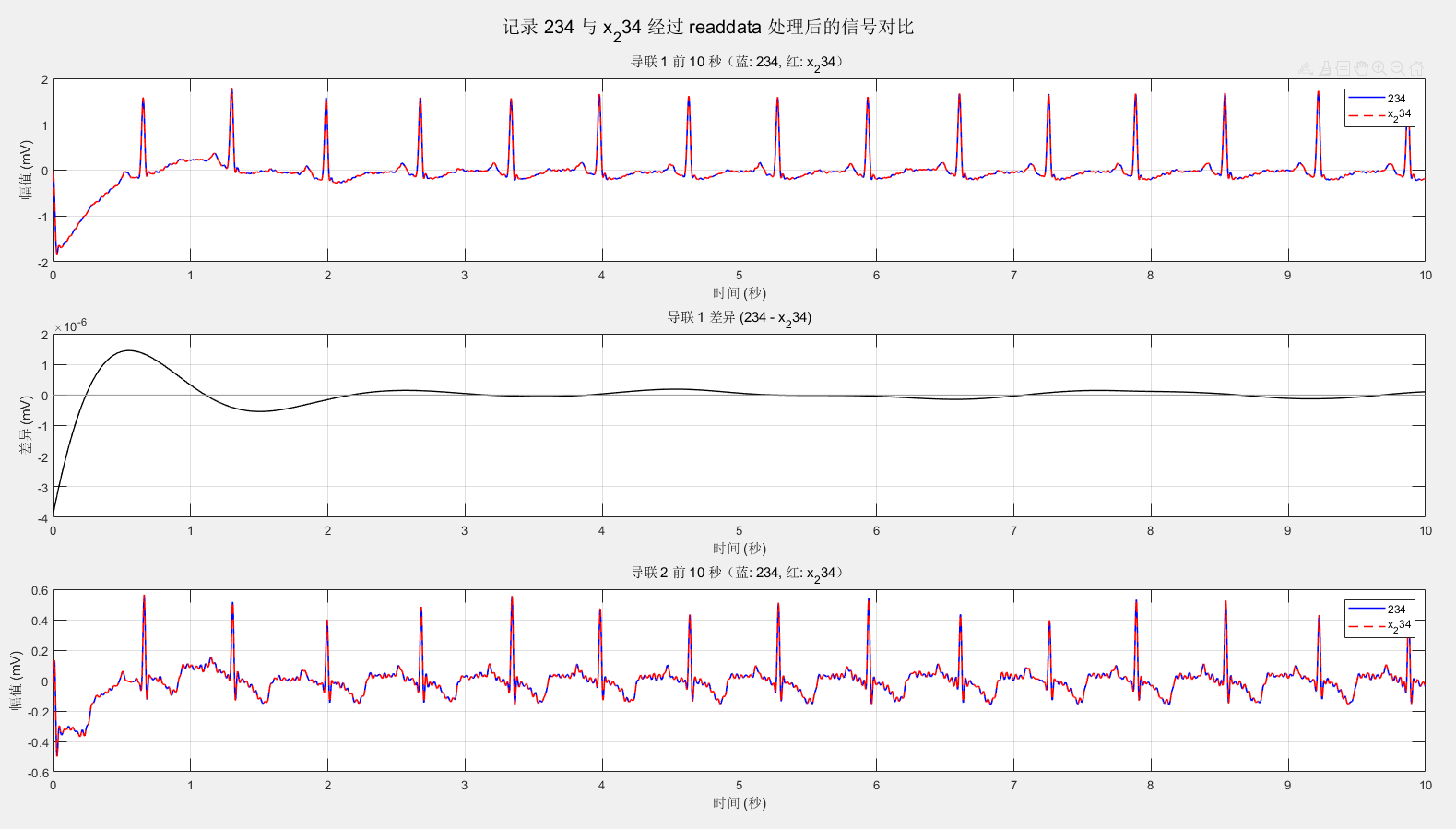
似乎没差别,我自己也看了其它信号并且放大范围,可以看出信号本身并无明显差异。
考虑到这里的问题主要是心律失常arrythmia,所以我们统计一下各个标签的数量:
Matlab
%% 总结各个文件中的注释标签类型及数量(基于 ASCII 字符映射)
clear; clc;
dataFolder = 'D:\ECG-Tran\mit-bih-arrhythmia-database';
recordNames = {
'100', '101', '102', '103', '104', '105', '106', '107', '108', '109', ...
'111', '112', '113', '114', '115', '116', '117', '118', '119', ...
'121', '122', '123', '124', ...
'200', '201', '202', '203', '205', '207', '208', '209', '210', ...
'212', '213', '214', '215', '217', '219', '220', '221', '222', '223', ...
'228', '230', '231', '232', '233', '234'
};
%% ========== 1. 构建映射表:字符 -> 描述 ==========
% 心搏类型 (Beat annotations)
beatMap = containers.Map();
beatMap('N') = '正常搏动';
beatMap('L') = '左束支传导阻滞搏动';
beatMap('R') = '右束支传导阻滞搏动';
beatMap('B') = '束支传导阻滞搏动(未指定)';
beatMap('A') = '房性早搏';
beatMap('a') = '异常房性早搏';
beatMap('J') = '交界性早搏';
beatMap('S') = '室上性早搏或异位搏动(房性或交界性)';
beatMap('V') = '室性早搏';
beatMap('r') = 'R-on-T 室性早搏';
beatMap('F') = '心室融合搏动(室性与正常融合)';
beatMap('e') = '房性逸搏';
beatMap('j') = '交界性逸搏';
beatMap('n') = '室上性逸搏(房性或交界性)';
beatMap('E') = '室性逸搏';
beatMap('/') = '起搏搏动';
beatMap('f') = '起搏与正常融合搏动';
beatMap('Q') = '不可分类搏动';
beatMap('?') = '学习期间未分类搏动';
% 非心搏注释 (Non-beat annotations)
nonBeatMap = containers.Map();
nonBeatMap('[') = '室扑/室颤开始';
nonBeatMap('!') = '室扑波';
nonBeatMap(']') = '室扑/室颤结束';
nonBeatMap('x') = '未传导P波 (阻滞的房性早搏)';
nonBeatMap('(') = '波形开始';
nonBeatMap(')') = '波形结束';
nonBeatMap('p') = 'P波峰值';
nonBeatMap('t') = 'T波峰值';
nonBeatMap('u') = 'U波峰值';
nonBeatMap('`') = 'PQ 连接处';
nonBeatMap(',') = 'J点';
nonBeatMap('~') = '(未夺获)起搏器伪迹 或 信号质量变化'; % 有两种用途
nonBeatMap('|') = '孤立的 QRS 样伪迹';
nonBeatMap('+') = '节律变化';
nonBeatMap('s') = 'ST段变化';
nonBeatMap('T') = 'T波变化';
nonBeatMap('*') = '收缩期';
nonBeatMap('D') = '舒张期';
nonBeatMap('=') = '测量注释';
nonBeatMap('"') = '评论注释';
nonBeatMap('@') = '指向外部数据的链接';
% 合并总映射(优先使用 beatMap,若不存在再查 nonBeatMap)
allMaps = containers.Map();
keys = [beatMap.keys, nonBeatMap.keys];
for i = 1:length(keys)
key = keys{i};
if beatMap.isKey(key)
allMaps(key) = beatMap(key);
elseif nonBeatMap.isKey(key)
allMaps(key) = nonBeatMap(key);
end
end
%% ========== 2. 处理每个记录 ==========
fprintf('%-8s | 标签类型及数量\n', '记录');
fprintf('%s\n', repmat('-', 100, 1));
for i = 1:length(recordNames)
recName = recordNames{i};
oldFolder = cd(dataFolder);
try
[~, atype] = rdann(recName, 'atr'); % atype 为数值数组,每个数值是 ASCII 码
catch
fprintf('%-8s | 无法读取注释\n', recName);
cd(oldFolder);
continue;
end
cd(oldFolder);
if isempty(atype)
fprintf('%-8s | 无注释\n', recName);
continue;
end
% 将 ASCII 码转换为字符,组成字符串数组
chars = cellfun(@(x) char(x), num2cell(atype), 'UniformOutput', false);
% 统计每个字符出现的次数
[uniqueChars, ~, idx] = unique(chars);
counts = accumarray(idx, 1);
% 生成输出字符串
infoParts = {};
for j = 1:length(uniqueChars)
ch = uniqueChars{j};
count = counts(j);
if allMaps.isKey(ch)
desc = allMaps(ch);
infoParts{end+1} = sprintf('''%s'' (%s): %d个', ch, desc, count);
else
% 未知字符:显示 ASCII 码和十六进制
asciiVal = double(ch);
infoParts{end+1} = sprintf('''%s'' (0x%02X, 未知): %d个', ch, asciiVal, count);
end
end
fprintf('%-8s | %s\n', recName, strjoin(infoParts, '; '));
end
Matlab
记录 | 标签类型及数量
----------------------------------------------------------------------------------------------------
100 | '+' (节律变化): 1个; 'A' (房性早搏): 33个; 'N' (正常搏动): 2239个; 'V' (室性早搏): 1个
101 | '+' (节律变化): 1个; 'A' (房性早搏): 3个; 'N' (正常搏动): 1860个; 'Q' (不可分类搏动): 2个; '|' (孤立的 QRS 样伪迹): 4个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 4个
102 | '+' (节律变化): 5个; '/' (起搏搏动): 2028个; 'N' (正常搏动): 99个; 'V' (室性早搏): 4个; 'f' (起搏与正常融合搏动): 56个
103 | '+' (节律变化): 1个; 'A' (房性早搏): 2个; 'N' (正常搏动): 2082个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 6个
104 | '+' (节律变化): 45个; '/' (起搏搏动): 1380个; 'N' (正常搏动): 163个; 'Q' (不可分类搏动): 18个; 'V' (室性早搏): 2个; 'f' (起搏与正常融合搏动): 666个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 37个
105 | '+' (节律变化): 1个; 'N' (正常搏动): 2526个; 'Q' (不可分类搏动): 5个; 'V' (室性早搏): 41个; '|' (孤立的 QRS 样伪迹): 30个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 88个
106 | '+' (节律变化): 41个; 'N' (正常搏动): 1507个; 'V' (室性早搏): 520个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 30个
107 | '+' (节律变化): 1个; '/' (起搏搏动): 2078个; 'V' (室性早搏): 59个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 2个
108 | '+' (节律变化): 1个; 'A' (房性早搏): 4个; 'F' (心室融合搏动(室性与正常融合)): 2个; 'N' (正常搏动): 1739个; 'V' (室性早搏): 17个; 'j' (交界性逸搏): 1个; 'x' (未传导P波 (阻滞的房性早搏)): 11个; '|' (孤立的 QRS 样伪迹): 8个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 41个
109 | '+' (节律变化): 1个; 'F' (心室融合搏动(室性与正常融合)): 2个; 'L' (左束支传导阻滞搏动): 2492个; 'V' (室性早搏): 38个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 2个
111 | '+' (节律变化): 1个; 'L' (左束支传导阻滞搏动): 2123个; 'V' (室性早搏): 1个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 8个
112 | '+' (节律变化): 1个; 'A' (房性早搏): 2个; 'N' (正常搏动): 2537个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 10个
113 | '+' (节律变化): 1个; 'N' (正常搏动): 1789个; 'a' (异常房性早搏): 6个
114 | '+' (节律变化): 3个; 'A' (房性早搏): 10个; 'F' (心室融合搏动(室性与正常融合)): 4个; 'J' (交界性早搏): 2个; 'N' (正常搏动): 1820个; 'V' (室性早搏): 43个; '|' (孤立的 QRS 样伪迹): 1个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 7个
115 | '+' (节律变化): 1个; 'N' (正常搏动): 1953个; '|' (孤立的 QRS 样伪迹): 6个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 2个
116 | '+' (节律变化): 1个; 'A' (房性早搏): 1个; 'N' (正常搏动): 2302个; 'V' (室性早搏): 109个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 8个
117 | '+' (节律变化): 1个; 'A' (房性早搏): 1个; 'N' (正常搏动): 1534个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 3个
118 | '+' (节律变化): 1个; 'A' (房性早搏): 96个; 'R' (右束支传导阻滞搏动): 2166个; 'V' (室性早搏): 16个; 'x' (未传导P波 (阻滞的房性早搏)): 10个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 12个
119 | '+' (节律变化): 103个; 'N' (正常搏动): 1543个; 'V' (室性早搏): 444个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 4个
121 | '+' (节律变化): 1个; 'A' (房性早搏): 1个; 'N' (正常搏动): 1861个; 'V' (室性早搏): 1个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 12个
122 | '+' (节律变化): 1个; 'N' (正常搏动): 2476个; '|' (孤立的 QRS 样伪迹): 2个
123 | '+' (节律变化): 1个; 'N' (正常搏动): 1515个; 'V' (室性早搏): 3个
124 | '+' (节律变化): 13个; 'A' (房性早搏): 2个; 'F' (心室融合搏动(室性与正常融合)): 5个; 'J' (交界性早搏): 29个; 'R' (右束支传导阻滞搏动): 1531个; 'V' (室性早搏): 47个; 'j' (交界性逸搏): 5个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 2个
200 | '+' (节律变化): 148个; 'A' (房性早搏): 30个; 'F' (心室融合搏动(室性与正常融合)): 2个; 'N' (正常搏动): 1743个; 'V' (室性早搏): 826个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 43个
201 | '+' (节律变化): 35个; 'A' (房性早搏): 30个; 'F' (心室融合搏动(室性与正常融合)): 2个; 'J' (交界性早搏): 1个; 'N' (正常搏动): 1625个; 'V' (室性早搏): 198个; 'a' (异常房性早搏): 97个; 'j' (交界性逸搏): 10个; 'x' (未传导P波 (阻滞的房性早搏)): 37个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 4个
202 | '+' (节律变化): 8个; 'A' (房性早搏): 36个; 'F' (心室融合搏动(室性与正常融合)): 1个; 'N' (正常搏动): 2061个; 'V' (室性早搏): 19个; 'a' (异常房性早搏): 19个; '|' (孤立的 QRS 样伪迹): 2个
203 | '+' (节律变化): 45个; 'F' (心室融合搏动(室性与正常融合)): 1个; 'N' (正常搏动): 2529个; 'Q' (不可分类搏动): 4个; 'V' (室性早搏): 444个; 'a' (异常房性早搏): 2个; '|' (孤立的 QRS 样伪迹): 26个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 57个
205 | '+' (节律变化): 13个; 'A' (房性早搏): 3个; 'F' (心室融合搏动(室性与正常融合)): 11个; 'N' (正常搏动): 2571个; 'V' (室性早搏): 71个; '|' (孤立的 QRS 样伪迹): 1个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 2个
207 | '!' (室扑波): 472个; '+' (节律变化): 24个; 'A' (房性早搏): 107个; 'E' (室性逸搏): 105个; 'L' (左束支传导阻滞搏动): 1457个; 'R' (右束支传导阻滞搏动): 86个; 'V' (室性早搏): 105个; '[' (室扑/室颤开始): 6个; ']' (室扑/室颤结束): 6个; '|' (孤立的 QRS 样伪迹): 2个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 15个
208 | '+' (节律变化): 53个; 'F' (心室融合搏动(室性与正常融合)): 373个; 'N' (正常搏动): 1586个; 'Q' (不可分类搏动): 2个; 'S' (室上性早搏或异位搏动(房性或交界性)): 2个; 'V' (室性早搏): 992个; '|' (孤立的 QRS 样伪迹): 8个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 24个
209 | '+' (节律变化): 21个; 'A' (房性早搏): 383个; 'N' (正常搏动): 2621个; 'V' (室性早搏): 1个; '|' (孤立的 QRS 样伪迹): 7个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 19个
210 | '+' (节律变化): 17个; 'E' (室性逸搏): 1个; 'F' (心室融合搏动(室性与正常融合)): 10个; 'N' (正常搏动): 2423个; 'V' (室性早搏): 194个; 'a' (异常房性早搏): 22个; '|' (孤立的 QRS 样伪迹): 1个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 17个
212 | '+' (节律变化): 1个; 'N' (正常搏动): 923个; 'R' (右束支传导阻滞搏动): 1825个; '|' (孤立的 QRS 样伪迹): 1个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 13个
213 | '+' (节律变化): 43个; 'A' (房性早搏): 25个; 'F' (心室融合搏动(室性与正常融合)): 362个; 'N' (正常搏动): 2641个; 'V' (室性早搏): 220个; 'a' (异常房性早搏): 3个
214 | '"' (评论注释): 1个; '+' (节律变化): 25个; 'F' (心室融合搏动(室性与正常融合)): 1个; 'L' (左束支传导阻滞搏动): 2003个; 'Q' (不可分类搏动): 2个; 'V' (室性早搏): 256个; '|' (孤立的 QRS 样伪迹): 5个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 4个
215 | '"' (评论注释): 2个; '+' (节律变化): 5个; 'A' (房性早搏): 3个; 'F' (心室融合搏动(室性与正常融合)): 1个; 'N' (正常搏动): 3195个; 'V' (室性早搏): 164个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 30个
217 | '+' (节律变化): 67个; '/' (起搏搏动): 1542个; 'N' (正常搏动): 244个; 'V' (室性早搏): 162个; 'f' (起搏与正常融合搏动): 260个; '|' (孤立的 QRS 样伪迹): 1个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 4个
219 | '"' (评论注释): 4个; '+' (节律变化): 21个; 'A' (房性早搏): 7个; 'F' (心室融合搏动(室性与正常融合)): 1个; 'N' (正常搏动): 2082个; 'V' (室性早搏): 64个; 'x' (未传导P波 (阻滞的房性早搏)): 133个
220 | '+' (节律变化): 16个; 'A' (房性早搏): 94个; 'N' (正常搏动): 1954个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 4个
221 | '+' (节律变化): 23个; 'N' (正常搏动): 2031个; 'V' (室性早搏): 396个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 12个
222 | '+' (节律变化): 136个; 'A' (房性早搏): 208个; 'J' (交界性早搏): 1个; 'N' (正常搏动): 2062个; 'j' (交界性逸搏): 212个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 15个
223 | '+' (节律变化): 28个; 'A' (房性早搏): 72个; 'F' (心室融合搏动(室性与正常融合)): 14个; 'N' (正常搏动): 2029个; 'V' (室性早搏): 473个; 'a' (异常房性早搏): 1个; 'e' (房性逸搏): 16个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 10个
228 | '"' (评论注释): 3个; '+' (节律变化): 41个; 'A' (房性早搏): 3个; 'N' (正常搏动): 1688个; 'V' (室性早搏): 362个; '|' (孤立的 QRS 样伪迹): 24个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 20个
230 | '+' (节律变化): 207个; 'N' (正常搏动): 2255个; 'V' (室性早搏): 1个; '|' (孤立的 QRS 样伪迹): 1个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 2个
231 | '"' (评论注释): 427个; '+' (节律变化): 11个; 'A' (房性早搏): 1个; 'N' (正常搏动): 314个; 'R' (右束支传导阻滞搏动): 1254个; 'V' (室性早搏): 2个; 'x' (未传导P波 (阻滞的房性早搏)): 2个
232 | '+' (节律变化): 1个; 'A' (房性早搏): 1382个; 'R' (右束支传导阻滞搏动): 397个; 'j' (交界性逸搏): 1个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 35个
233 | '+' (节律变化): 71个; 'A' (房性早搏): 7个; 'F' (心室融合搏动(室性与正常融合)): 11个; 'N' (正常搏动): 2230个; 'V' (室性早搏): 831个; '|' (孤立的 QRS 样伪迹): 2个
234 | '+' (节律变化): 3个; 'J' (交界性早搏): 50个; 'N' (正常搏动): 2700个; 'V' (室性早搏): 3个; '~' ((未夺获)起搏器伪迹 或 信号质量变化): 8个读取数据与降采样,该数据库是360Hz,为了配合我们之前的数据,我们可以选择将其降采样(需要将注释同步考虑):
Matlab
function [sig, Fs, tm, annot_pos, annot_sym] = readdata(recordName, dataFolder, targetFs)
% 读取心电图信号,从第5秒开始裁切,进行去趋势和带通滤波,并降采样至目标频率
% 同时读取 .atr 注释,并将注释位置同步映射到降采样后的时间轴上
% 输入:
% recordName - 记录名称
% dataFolder - 数据文件夹路径
% targetFs - 目标采样率 (Hz),默认 128
% 输出:
% sig - 处理后的双导联信号 [nSamples × 2]
% Fs - 目标采样率 (Hz)
% tm - 时间向量(从0开始,对应降采样后的信号)
% annot_pos - 降采样后的注释样本索引 (列向量,与 annot_sym 对应)
% annot_sym - 注释符号的 cell 数组 (如 {'N','V','A',...})
if nargin < 3
targetFs = 128; % 默认目标采样率 128 Hz
end
oldFolder = cd(dataFolder);
cleanup = onCleanup(@() cd(oldFolder));
% ------------------ 读取信号 ------------------
[sig_full, origFs] = rdsamp(recordName);
if isempty(sig_full)
error('无法读取记录 %s 的信号', recordName);
end
% 第5秒对应的原始样本起始索引 (1-based)
start_second = 5;
startSample = round(start_second * origFs) + 1;
if startSample > size(sig_full, 1)
error('第%d秒(样本 %d)超出信号长度(%d)', start_second, startSample, size(sig_full,1));
end
% 裁切信号
sig_cropped = sig_full(startSample:end, :);
fprintf('记录 %s:从第%d秒(样本 %d)开始裁切,剩余 %d 个样本\n', ...
recordName, start_second, startSample, size(sig_cropped,1));
% 去趋势 + 带通滤波 (0.5-40 Hz)
sig_filtered = zeros(size(sig_cropped));
nyquist = origFs / 2;
[b, a] = butter(4, [0.5/nyquist, 40/nyquist], 'bandpass');
for lead = 1:2
signal_detrend = detrend(sig_cropped(:, lead));
sig_filtered(:, lead) = filtfilt(b, a, signal_detrend);
end
% 降采样至目标采样率
if origFs ~= targetFs
[p, q] = rat(targetFs / origFs, 1e-12);
fprintf('重采样:从 %.1f Hz 到 %.1f Hz (因子 %.6f)\n', origFs, targetFs, p/q);
n_out = ceil(size(sig_filtered, 1) * p / q);
sig = zeros(n_out, 2);
for lead = 1:2
sig(:, lead) = resample(sig_filtered(:, lead), p, q);
end
Fs = targetFs;
else
sig = sig_filtered;
Fs = origFs;
end
% 时间向量 (秒,从0开始)
tm = (0:size(sig,1)-1) / Fs;
% ------------------ 读取并同步注释 ------------------
annot_pos = [];
annot_sym = {};
try
% 读取 atr 注释:ann_orig 为样本位置(0-based),anntype_orig 为 ASCII 码数值
[ann_orig, anntype_orig] = rdann(recordName, 'atr');
if isempty(ann_orig)
fprintf('记录 %s 无注释数据\n', recordName);
return;
end
% 转换为 1-based 索引,并转换 ASCII 码为字符
orig_idx = ann_orig + 1; % 转为 1-based
sym_cell = cellfun(@(x) char(x), num2cell(anntype_orig), 'UniformOutput', false);
% 只保留裁切起始点之后的注释
keep = (orig_idx >= startSample);
orig_idx = orig_idx(keep);
sym_cell = sym_cell(keep);
if isempty(orig_idx)
fprintf('记录 %s 在第5秒后无注释\n', recordName);
return;
end
% 计算裁切后的相对样本索引 (基于原始采样率)
cropped_idx = orig_idx - startSample + 1; % 1-based,相对于 sig_cropped
% 裁切后的时间 (秒)
t_cropped = (cropped_idx - 1) / origFs;
% 映射到降采样后的样本索引 (四舍五入)
new_idx = round(t_cropped * targetFs) + 1;
% 剔除越界的索引
valid = (new_idx >= 1) & (new_idx <= size(sig,1));
annot_pos = new_idx(valid);
annot_sym = sym_cell(valid);
% 可选:去除重复位置上的多余注释(保留第一个)
[annot_pos, unique_ia] = unique(annot_pos, 'stable');
annot_sym = annot_sym(unique_ia);
fprintf('成功同步 %d 个注释到降采样信号 (总注释 %d)\n', length(annot_pos), sum(keep));
catch ME
warning('读取或同步注释时出错: %s', ME.message);
end
end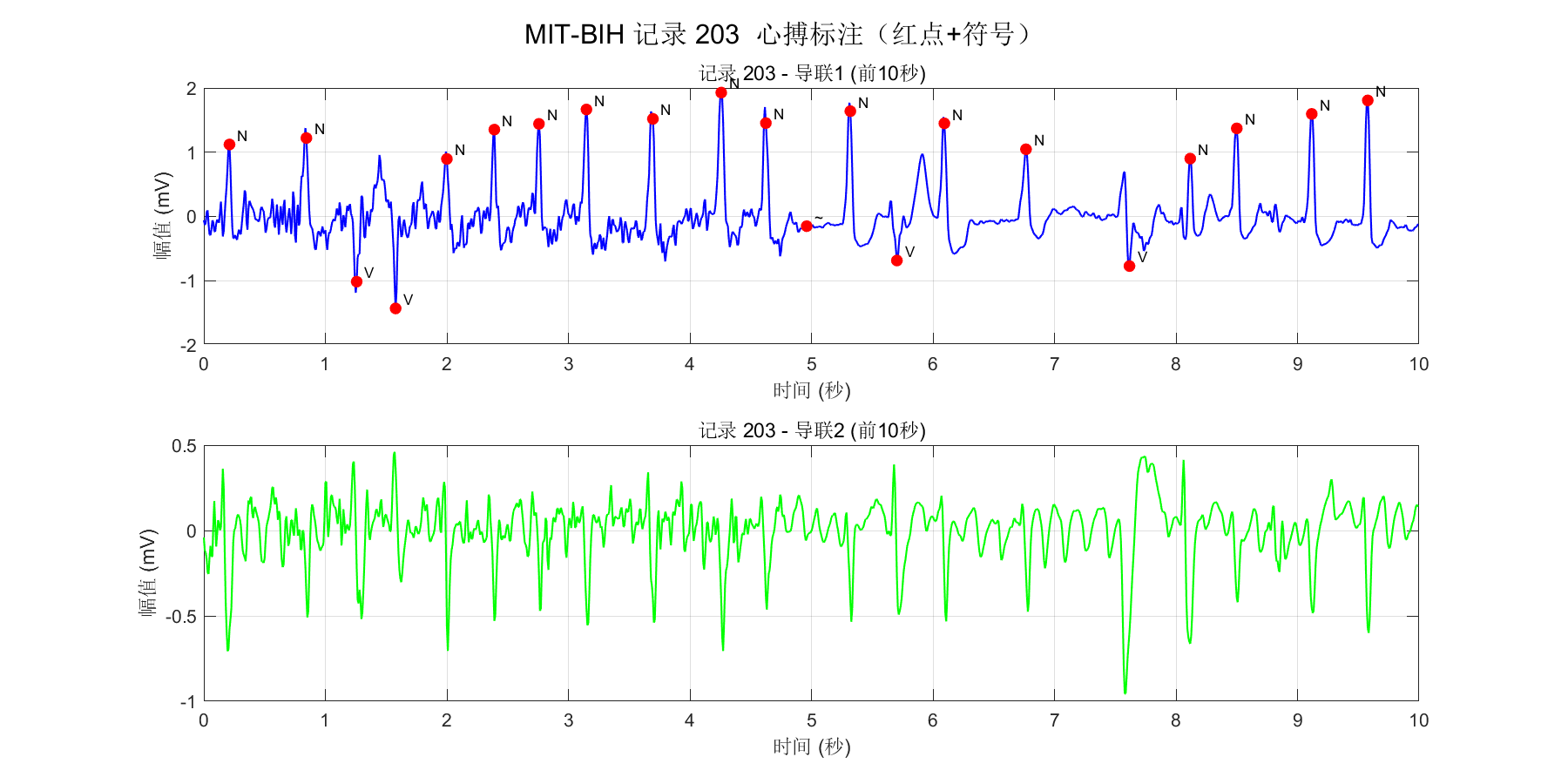
各种标签中表示R峰位置的有:
Matlab
beat_chars = {'N','L','R','B','A','a','J','S','V','r','F','e','j','n','E','/','f','Q','?'};我在这里写一个输出R峰索引的脚本:
Matlab
function [r_idx, r_times] = get_r_peaks(annot_pos, annot_sym, Fs)
% GET_R_PEAKS 从注释中筛选出指定心搏类型的R波位置
% 输入:
% annot_pos - 降采样后的注释样本索引 (列向量)
% annot_sym - 注释符号的 cell 数组 (与 annot_pos 对应)
% Fs - 当前信号的采样率 (Hz)
% 输出:
% r_idx - 筛选后的R波样本索引 (列向量)
% r_times - 对应的时刻 (秒,列向量)
% 定义需要保留的心搏类型
beat_chars = {'N','L','R','B','A','a','J','S','V','r','F','e','j','n','E','/','f','Q','?'};
% 检查输入是否为空
if isempty(annot_pos) || isempty(annot_sym)
r_idx = [];
r_times = [];
fprintf('警告:没有提供任何注释,无法提取R波。\n');
return;
end
% 确保 annot_pos 是列向量
annot_pos = annot_pos(:);
% 确保 annot_sym 是列 cell 数组
if isrow(annot_sym)
annot_sym = annot_sym';
end
% 创建逻辑掩码:每个符号是否属于目标心搏类型
keep_mask = ismember(annot_sym, beat_chars);
% 筛选出R波对应的样本索引
r_idx = annot_pos(keep_mask);
% 计算对应的时间 (秒) tm = (idx-1)/Fs
r_times = (r_idx - 1) / Fs;
end**********END**********