📘 PySpark 核心实操入门指南:从数据输入到输出,全面掌握 Spark 计算范式
本文目标
本文以 真实案例 + 深度拆解 的方式,带你从零开始掌握 PySpark 的核心机制,涵盖:
- RDD 基本概念与创建
- 数据类型转换:
T → T与T → U - 核心算子详解(
map,flatMap,filter,distinct,sortBy,reduceByKey) - 如何修改 RDD 分区?
- 如何修复
MSVCR120.dll缺失?(Windows 专属) - 如何解决 Spark 超时问题?
- 两个实战案例:城市销量排名 & 词频统计
一、PySpark 环境搭建基础配置(关键!)
特此强调:Windows 用户务必设置以下环境变量,否则几乎所有操作都可能失败!
python
import os
# 1. 指定 Python 解释器路径(必须!)
os.environ["PYSPARK_PYTHON"] = "D:\\APP\\Anaconda\\envs\\spark_env\\python.exe"
# 2. Windows 超时问题(防止 task 超时崩溃)
os.environ['PYSPARK_TIMEOUT'] = '600'
os.environ['PYSPARK_DRIVER_TIMEOUT'] = '600'
# 3. Hadoop native dll 加载导致 MSVCR120.dll 报错
os.environ['PATH'] += os.pathsep + 'E:\\APP\\hadoop-3.4.2\\bin'
# 4. 强制设置 driver python(可选但推荐)
os.environ["PYSPARK_DRIVER_PYTHON"] = "D:\\APP\\Anaconda\\envs\\spark_env\\python.exe"为什么这步如此重要?
MSVCR120.dll是 Visual C++ 2013 运行时库 ,常在 Hadoop 的hadoop.dll调用失败时出现。- 即使你不用
saveAsTextFile(),如果PATH中包含hadoop/bin,Spark 仍会触发 native IO!- 终极建议:仅当必须使用原生 IO 时才保留
PATH;否则删除该路径!
如何检查到是 MSVCR120.dll 的问题: 这是我的报错
bash
from pyspark import SparkConf,SparkContext
import os
os.environ["PYSPARK_PYTHON"] = "D:\APP\Anaconda\envs\spark_env\python.exe"
os.environ['PATH'] += os.pathsep + 'E:\\APP\\hadoop-3.4.2\\bin'
conf = SparkConf().setMaster("local").setAppName("test_spark_app")
sc = SparkContext(conf=conf)
rdd = sc.parallelize([1,2,3,4,5])
rdd.saveAsTextFile("D:/output1")
sc.stop()D:\APP\Anaconda\envs\spark_env\python.exe D:\PythonProjects\python\day11_PySpark\数据输出\输出为文本文档.py
Setting default log level to "WARN".
To adjust logging level use sc.setLogLevel(newLevel). For SparkR, use setLogLevel(newLevel).
26/04/01 22:22:55 WARN NativeCodeLoader: Unable to load native-hadoop library for your platform... using builtin-java classes where applicable
26/04/01 22:22:58 ERROR SparkHadoopWriter: Aborting job job_202604012222567528972640969878075_0003.
java.lang.UnsatisfiedLinkError: org.apache.hadoop.io.nativeio.NativeIO$Windows.access0(Ljava/lang/String;I)Z
at org.apache.hadoop.io.nativeio.NativeIO$Windows.access0(Native Method)
at org.apache.hadoop.io.nativeio.NativeIO$Windows.access(NativeIO.java:793)
at org.apache.hadoop.fs.FileUtil.canRead(FileUtil.java:1249)
at org.apache.hadoop.fs.FileUtil.list(FileUtil.java:1454)
at org.apache.hadoop.fs.RawLocalFileSystem.listStatus(RawLocalFileSystem.java:601)
at org.apache.hadoop.fs.FileSystem.listStatus(FileSystem.java:1972)
at org.apache.hadoop.fs.FileSystem.listStatus(FileSystem.java:2014)
at org.apache.hadoop.fs.ChecksumFileSystem.listStatus(ChecksumFileSystem.java:761)
at org.apache.hadoop.fs.FileSystem.listStatus(FileSystem.java:1972)
at org.apache.hadoop.fs.FileSystem.listStatus(FileSystem.java:2014)
at org.apache.hadoop.mapreduce.lib.output.FileOutputCommitter.getAllCommittedTaskPaths(FileOutputCommitter.java:334)
at org.apache.hadoop.mapreduce.lib.output.FileOutputCommitter.commitJobInternal(FileOutputCommitter.java:404)
at org.apache.hadoop.mapreduce.lib.output.FileOutputCommitter.commitJob(FileOutputCommitter.java:377)
at org.apache.hadoop.mapred.FileOutputCommitter.commitJob(FileOutputCommitter.java:136)
at org.apache.hadoop.mapred.OutputCommitter.commitJob(OutputCommitter.java:291)
at org.apache.spark.internal.io.HadoopMapReduceCommitProtocol.commitJob(HadoopMapReduceCommitProtocol.scala:192)
at org.apache.spark.internal.io.SparkHadoopWriter$.$anonfun$write$3(SparkHadoopWriter.scala:100)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.util.Utils$.timeTakenMs(Utils.scala:552)
at org.apache.spark.internal.io.SparkHadoopWriter$.write(SparkHadoopWriter.scala:100)
at org.apache.spark.rdd.PairRDDFunctions.$anonfun$saveAsHadoopDataset$1(PairRDDFunctions.scala:1091)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.PairRDDFunctions.saveAsHadoopDataset(PairRDDFunctions.scala:1089)
at org.apache.spark.rdd.PairRDDFunctions.$anonfun$saveAsHadoopFile$4(PairRDDFunctions.scala:1062)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.PairRDDFunctions.saveAsHadoopFile(PairRDDFunctions.scala:1027)
at org.apache.spark.rdd.PairRDDFunctions.$anonfun$saveAsHadoopFile$3(PairRDDFunctions.scala:1009)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.PairRDDFunctions.saveAsHadoopFile(PairRDDFunctions.scala:1008)
at org.apache.spark.rdd.PairRDDFunctions.$anonfun$saveAsHadoopFile$2(PairRDDFunctions.scala:965)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.PairRDDFunctions.saveAsHadoopFile(PairRDDFunctions.scala:963)
at org.apache.spark.rdd.RDD.$anonfun$saveAsTextFile$2(RDD.scala:1620)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.RDD.saveAsTextFile(RDD.scala:1620)
at org.apache.spark.rdd.RDD.$anonfun$saveAsTextFile$1(RDD.scala:1606)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.RDD.saveAsTextFile(RDD.scala:1606)
at org.apache.spark.api.java.JavaRDDLike.saveAsTextFile(JavaRDDLike.scala:564)
at org.apache.spark.api.java.JavaRDDLike.saveAsTextFile$(JavaRDDLike.scala:563)
at org.apache.spark.api.java.AbstractJavaRDDLike.saveAsTextFile(JavaRDDLike.scala:45)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(Unknown Source)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(Unknown Source)
at java.lang.reflect.Method.invoke(Unknown Source)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:374)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.ClientServerConnection.waitForCommands(ClientServerConnection.java:182)
at py4j.ClientServerConnection.run(ClientServerConnection.java:106)
at java.lang.Thread.run(Unknown Source)
Traceback (most recent call last):
File "D:\PythonProjects\python\day11_PySpark\数据输出\输出为文本文档.py", line 9, in <module>
rdd.saveAsTextFile("D:/output1")
File "D:\APP\Anaconda\envs\spark_env\lib\site-packages\pyspark\rdd.py", line 3425, in saveAsTextFile
keyed._jrdd.map(self.ctx._jvm.BytesToString()).saveAsTextFile(path)
File "D:\APP\Anaconda\envs\spark_env\lib\site-packages\py4j\java_gateway.py", line 1322, in __call__
return_value = get_return_value(
File "D:\APP\Anaconda\envs\spark_env\lib\site-packages\py4j\protocol.py", line 326, in get_return_value
raise Py4JJavaError(
py4j.protocol.Py4JJavaError: An error occurred while calling o33.saveAsTextFile.
: org.apache.spark.SparkException: Job aborted.
at org.apache.spark.internal.io.SparkHadoopWriter$.write(SparkHadoopWriter.scala:106)
at org.apache.spark.rdd.PairRDDFunctions.$anonfun$saveAsHadoopDataset$1(PairRDDFunctions.scala:1091)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.PairRDDFunctions.saveAsHadoopDataset(PairRDDFunctions.scala:1089)
at org.apache.spark.rdd.PairRDDFunctions.$anonfun$saveAsHadoopFile$4(PairRDDFunctions.scala:1062)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.PairRDDFunctions.saveAsHadoopFile(PairRDDFunctions.scala:1027)
at org.apache.spark.rdd.PairRDDFunctions.$anonfun$saveAsHadoopFile$3(PairRDDFunctions.scala:1009)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.PairRDDFunctions.saveAsHadoopFile(PairRDDFunctions.scala:1008)
at org.apache.spark.rdd.PairRDDFunctions.$anonfun$saveAsHadoopFile$2(PairRDDFunctions.scala:965)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.PairRDDFunctions.saveAsHadoopFile(PairRDDFunctions.scala:963)
at org.apache.spark.rdd.RDD.$anonfun$saveAsTextFile$2(RDD.scala:1620)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.RDD.saveAsTextFile(RDD.scala:1620)
at org.apache.spark.rdd.RDD.$anonfun$saveAsTextFile$1(RDD.scala:1606)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:151)
at org.apache.spark.rdd.RDDOperationScope$.withScope(RDDOperationScope.scala:112)
at org.apache.spark.rdd.RDD.withScope(RDD.scala:407)
at org.apache.spark.rdd.RDD.saveAsTextFile(RDD.scala:1606)
at org.apache.spark.api.java.JavaRDDLike.saveAsTextFile(JavaRDDLike.scala:564)
at org.apache.spark.api.java.JavaRDDLike.saveAsTextFile$(JavaRDDLike.scala:563)
at org.apache.spark.api.java.AbstractJavaRDDLike.saveAsTextFile(JavaRDDLike.scala:45)
at sun.reflect.NativeMethodAccessorImpl.invoke0(Native Method)
at sun.reflect.NativeMethodAccessorImpl.invoke(Unknown Source)
at sun.reflect.DelegatingMethodAccessorImpl.invoke(Unknown Source)
at java.lang.reflect.Method.invoke(Unknown Source)
at py4j.reflection.MethodInvoker.invoke(MethodInvoker.java:244)
at py4j.reflection.ReflectionEngine.invoke(ReflectionEngine.java:374)
at py4j.Gateway.invoke(Gateway.java:282)
at py4j.commands.AbstractCommand.invokeMethod(AbstractCommand.java:132)
at py4j.commands.CallCommand.execute(CallCommand.java:79)
at py4j.ClientServerConnection.waitForCommands(ClientServerConnection.java:182)
at py4j.ClientServerConnection.run(ClientServerConnection.java:106)
at java.lang.Thread.run(Unknown Source)
Caused by: java.lang.UnsatisfiedLinkError: org.apache.hadoop.io.nativeio.NativeIO$Windows.access0(Ljava/lang/String;I)Z
at org.apache.hadoop.io.nativeio.NativeIO$Windows.access0(Native Method)
at org.apache.hadoop.io.nativeio.NativeIO$Windows.access(NativeIO.java:793)
at org.apache.hadoop.fs.FileUtil.canRead(FileUtil.java:1249)
at org.apache.hadoop.fs.FileUtil.list(FileUtil.java:1454)
at org.apache.hadoop.fs.RawLocalFileSystem.listStatus(RawLocalFileSystem.java:601)
at org.apache.hadoop.fs.FileSystem.listStatus(FileSystem.java:1972)
at org.apache.hadoop.fs.FileSystem.listStatus(FileSystem.java:2014)
at org.apache.hadoop.fs.ChecksumFileSystem.listStatus(ChecksumFileSystem.java:761)
at org.apache.hadoop.fs.FileSystem.listStatus(FileSystem.java:1972)
at org.apache.hadoop.fs.FileSystem.listStatus(FileSystem.java:2014)
at org.apache.hadoop.mapreduce.lib.output.FileOutputCommitter.getAllCommittedTaskPaths(FileOutputCommitter.java:334)
at org.apache.hadoop.mapreduce.lib.output.FileOutputCommitter.commitJobInternal(FileOutputCommitter.java:404)
at org.apache.hadoop.mapreduce.lib.output.FileOutputCommitter.commitJob(FileOutputCommitter.java:377)
at org.apache.hadoop.mapred.FileOutputCommitter.commitJob(FileOutputCommitter.java:136)
at org.apache.hadoop.mapred.OutputCommitter.commitJob(OutputCommitter.java:291)
at org.apache.spark.internal.io.HadoopMapReduceCommitProtocol.commitJob(HadoopMapReduceCommitProtocol.scala:192)
at org.apache.spark.internal.io.SparkHadoopWriter$.$anonfun$write$3(SparkHadoopWriter.scala:100)
at scala.runtime.java8.JFunction0$mcV$sp.apply(JFunction0$mcV$sp.java:23)
at org.apache.spark.util.Utils$.timeTakenMs(Utils.scala:552)
at org.apache.spark.internal.io.SparkHadoopWriter$.write(SparkHadoopWriter.scala:100)
... 51 more
进程已结束,退出代码为 1由于找不到 MSVCR120.dll,无法继续执行代码。重新安装程序可能会解决此问题。 是 Windows 系统缺少 C++ 运行时库 的典型问题。
🚨 根本原因:
MSVCR120.dll 是 Microsoft Visual C++ 2013 Redistributable 的一部分。
它被 Hadoop 的原生 .dll 文件(如 hadoop.dll)所依赖,当你使用 saveAsTextFile() 时,Spark 会尝试加载 Hadoop 的本地库,而这些库又依赖于 VC++ 运行时。
即使你现在不用 saveAsTextFile(),你之前配置了 PATH += E:\APP\hadoop-3.4.2\bin,说明你还在用 Hadoop 二进制文件,所以问题依然存在。
我一开始怀疑是缺少winutils 环境配置 winutils配置可以看我上个文章 或者是 winutils.exe 权限不对
你的 winutils.exe 必须放在hadoop的bin目录下 E:\APP\hadoop-3.4.2\bin\winutils.exe
然后检查 权限
以 管理员身份 打开 CMD
一定要右键 → 以管理员身份运行命令提示符
. 执行你这条命令
bash
```bash
winutils.exe chmod 777 C:\tmp\Hive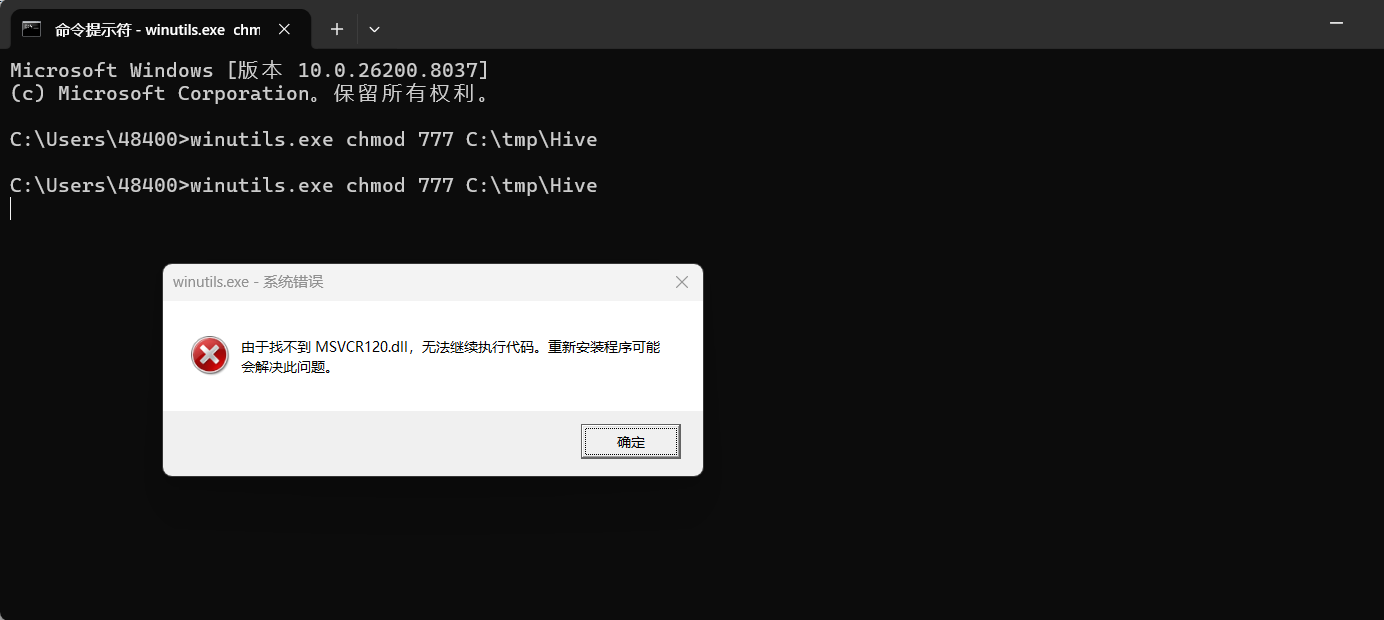
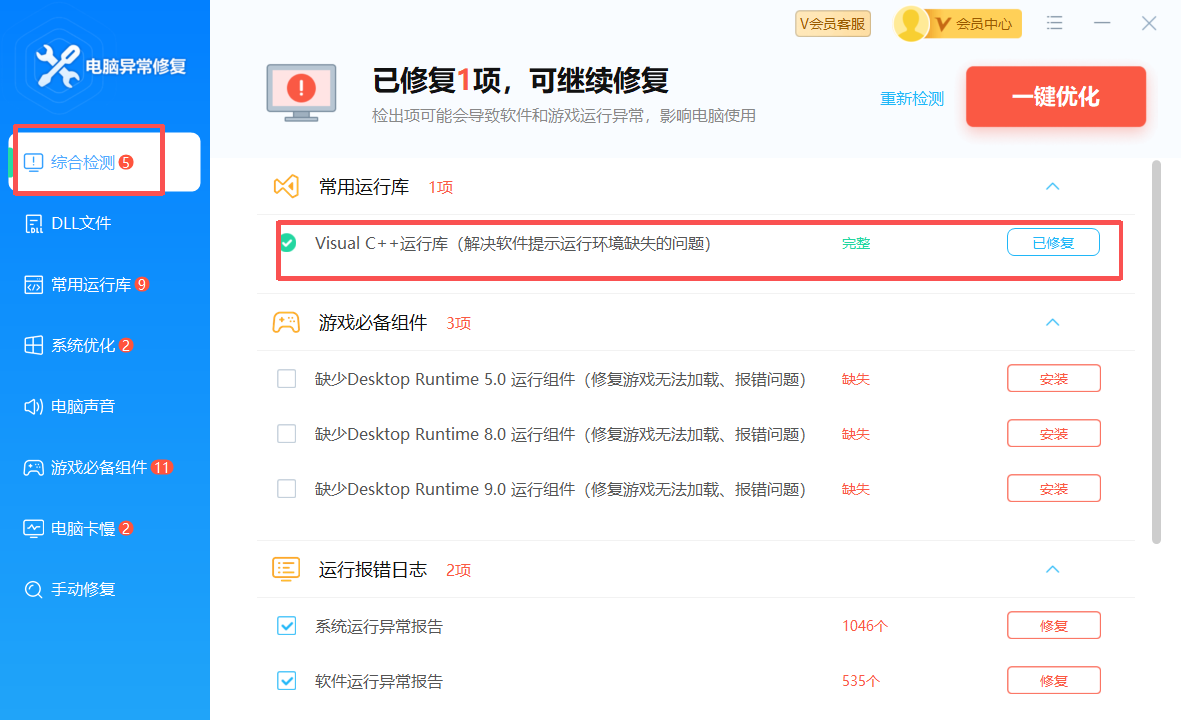
可以用联想的电脑异常修复直接修复
📌 二、RDD 基础概念与创建方式 💡
什么是 RDD?
RDD(Resilient Distributed Dataset):弹性分布式数据集,是 Spark 的核心抽象。它是一个不可变、可并行、容错的数据集合。
创建 RDD 的 6 种方式:
| 类型 | 示例 | 说明 |
|---|---|---|
| Python 列表 | sc.parallelize([1,2,3]) |
最常用 |
| 字典 | sc.parallelize({"k":1}) |
返回 [{"k":1}] |
| 元组 | sc.parallelize(("a","b")) |
返回 [("a","b")] |
| 集合 | sc.parallelize([1,1,2,3]) |
包含重复项 |
| 字符串 | sc.parallelize("abc") |
每个字符为一个元素 |
| 文件 | sc.textFile("path.txt") |
逐行读取文本 |
推荐使用
parallelize()+textFile()搭配处理本地数据。
三、核心算子详解:类型转换与功能解析
1. map(T) → U:传入类型与返回类型不一致
作用:对每个元素进行函数映射,返回新值
python
rdd = sc.parallelize([1,2,3,4,5])
rdd2 = rdd.map(lambda x: x * 2)
print(rdd2.collect()) # 输出: [2, 4, 6, 8, 10]| 特点 | 说明 |
|---|---|
T → U |
输入是 int,输出是 int(类型一致),但语义上是"变换" |
| 按元素处理 | 每个元素独立转换 |
| 无聚合 | 不改变数据总量 |
适用场景:数据清洗、类型转换、数学计算
2. map(T) → T:传入类型与返回类型一致
作用:返回原类型,但可以修改内容
python
rdd = sc.parallelize(["apple", "banana"])
rdd2 = rdd.map(lambda x: x.upper())
print(rdd2.collect()) # ['APPLE', 'BANANA']虽然返回类型仍是
str,但内容被修改 → 仍属于T → T
适用场景:字符串大小写转换、去除空格、文本标准化
3. flatMap(T) → U:扁平化处理,去嵌套
作用:将结果展开成平面列表
python
rdd = sc.parallelize(["hello world", "py spark"])
rdd2 = rdd.flatMap(lambda x: x.split(" "))
print(rdd2.collect()) # ['hello', 'world', 'py', 'spark']| 区别 | map vs flatMap |
|---|---|
map 返回 [[x], [y]] |
返回嵌套列表 |
flatMap 返回 [x, y] |
展开为单层列表 |
适用场景:文本分词、多行拆分、列表合并
4. filter(T) → T:条件过滤
作用:根据返回
True/False过滤元素
python
rdd = sc.parallelize([1,2,3,4,5,6])
rdd2 = rdd.filter(lambda x: x % 2 == 0)
print(rdd2.collect()) # [2, 4, 6]适用场景:筛选城市、过滤商品类别、去除空值
5. distinct():去重
作用:返回去重后的集合(全局去重)
python
rdd = sc.parallelize([1,2,2,3,4,4,5])
rdd2 = rdd.distinct()
print(rdd2.collect()) # [1, 2, 3, 4, 5]注意:distinct() 耗时高,会 Shuffle 所有数据!慎用于大数据集!
6. sortBy(keyfunc, ascending):排序
作用:按指定规则排序
python
rdd = sc.parallelize([1,5,3,2,4])
rdd2 = rdd.sortBy(lambda x: x, ascending=False)
print(rdd2.collect()) # [5, 4, 3, 2, 1]适用场景:销售排名、分数排序、Top N 推荐
7. reduceByKey(func):分组聚合(键值对专属)
前提:RDD 数据必须是
(K, V)元组形式
python
rdd = sc.parallelize([("a",1),("a",2),("b",3),("b",4)])
rdd2 = rdd.reduceByKey(lambda a, b: a + b)
print(rdd2.collect()) # [('a', 3), ('b', 7)]| 要求 | 说明 |
|---|---|
必须是 (K, V) |
否则报错 |
func(V,V) → V |
两个 value 聚合出一个 value |
| 本地预聚合 | 减少网络传输,效率极高 |
适用场景:统计商品销量、城市销售额、词频统计
四、修改 RDD 分区的 3 种方法
| 方法 | 说明 | 代码示例 |
|---|---|---|
set("spark.default.parallelism", "1") |
设置全局并行度 | conf.set("spark.default.parallelism", "1") |
numSlices=1 |
parallelize 时指定分区数 |
sc.parallelize([1,2,3], 1) |
repartition(n) |
重新分区(常用于调优) | rdd.repartition(2) |
建议 :小数据集用
numSlices=1避免创建过多任务。
五、重难点攻克:如何修复 MSVCR120.dll 缺失问题?💥
报错信息:
由于找不到 MSVCR120.dll,无法继续执行代码
🔍 根本原因:
- Spark 调用
hadoop.dll会依赖 VC++ 2013 运行时。 PATH中包含E:\APP\hadoop-3.4.2\bin→ 启动 native IO → 加载 DLL → 找不到函数 → 报错。
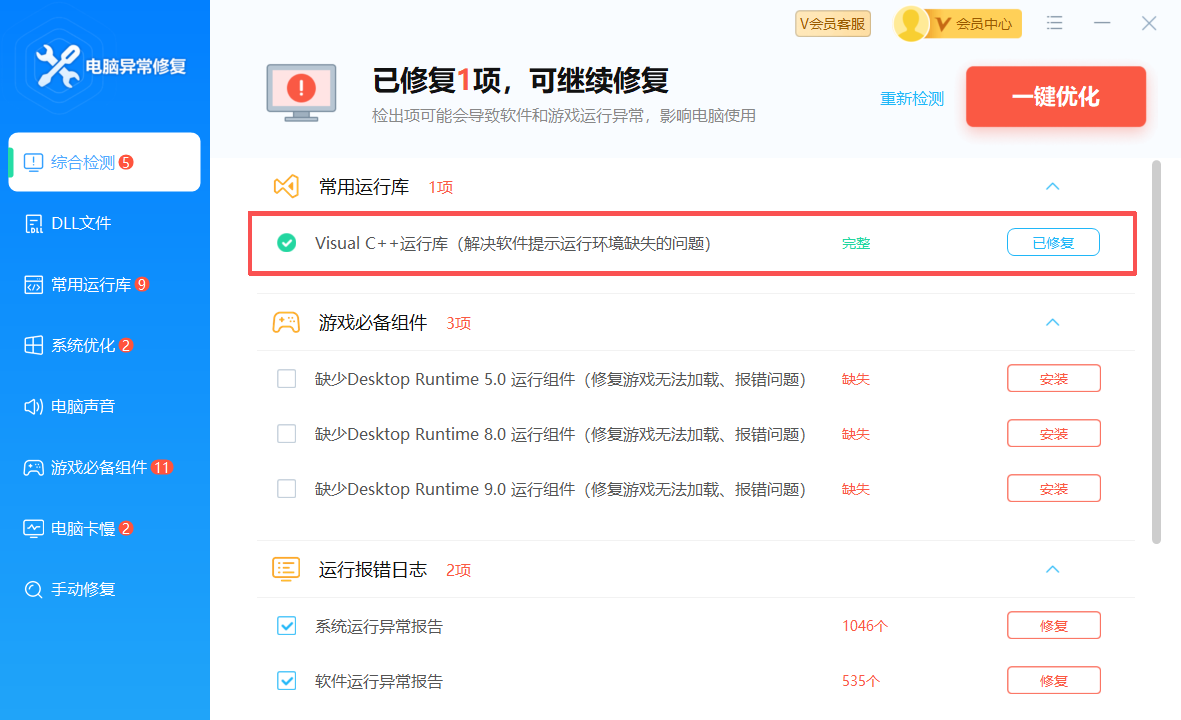
终极解决方案:
修复 MSVCR120.dll 可以用联想的电脑异常修复直接修复
六、案例实战:综合数据分析
案例 1:城市销量排名(P09_案例2.py)
需求:
- 读取
sales.txt文件(JSON 格式) - 统计每个城市销售额从大到小排序
- 获取全部城市商品类别(去重)
- 查询"北京"有哪些商品类别(过滤 + 去重)
步骤详解:
python
import json
from pyspark import SparkContext, SparkConf
import os
os.environ["PYSPARK_PYTHON"] = "D:\\APP\\Anaconda\\envs\\spark_env\\python.exe"
os.environ['PYSPARK_TIMEOUT'] = '600'
os.environ['PYSPARK_DRIVER_TIMEOUT'] = '600'
conf = SparkConf().setMaster("local[*]").setAppName("sales_analysis")
sc = SparkContext(conf=conf)
# 1. 读取文件并转为字典
file_rdd = sc.textFile("D:\\python_testdata\\sales.txt")
dict_rdd = file_rdd.map(lambda x: json.loads(x.rstrip().rstrip(',')))
# 2. 各城市销售额排名
city_money = dict_rdd.map(lambda x: (x['areaName'], int(x['money'])))
city_total = city_money.reduceByKey(lambda a, b: a + b)
city_rank = city_total.sortBy(lambda x: x[1], ascending=False)
print("各城市销售额排名:")
print(city_rank.collect())
# 3. 全部城市有哪些商品类别(去重)
categories = dict_rdd.map(lambda x: x['category'])
unique_categories = categories.distinct()
print("全部商品类别:")
print(unique_categories.collect())
# 4. 北京市的商品类别
beijing_rdd = dict_rdd.filter(lambda x: x['areaName'] == '北京')
beijing_cats = beijing_rdd.map(lambda x: x['category']).distinct()
print("北京市售卖的商品类别:")
print(beijing_cats.collect())
sc.stop()知识点总结:
json.loads():解析 JSON 字符串map:提取字段reduceByKey:聚合销售额sortBy:排序filter+distinct:组合过滤去重
案例 2:词频统计(P05_PySpark案例.py)
需求:统计 words.txt 中每个词出现次数
python
from pyspark import SparkContext, SparkConf
import os
os.environ["PYSPARK_PYTHON"] = "D:\\APP\\Anaconda\\envs\\spark_env\\python.exe"
os.environ['PYSPARK_TIMEOUT'] = '600'
conf = SparkConf().setMaster("local[*]").setAppName("word_count")
sc = SparkContext(conf=conf)
rdd = sc.textFile("D:\\python_testdata\\words.txt")
words = rdd.flatMap(lambda x: x.split(" "))
word_pairs = words.map(lambda x: (x, 1))
word_count = word_pairs.reduceByKey(lambda a, b: a + b)
print("词频统计结果:")
print(word_count.collect())
sc.stop()知识点:
flatMap分词map→(word, 1)reduceByKey聚合- 典型的 MapReduce 模型
七、总结与建议
| 项目 | 推荐做法 |
|---|---|
| 数据输出 | ✅ 使用 collect() + open() 写文件 ,避免 saveAsTextFile() |
| 环境配置 | ✅ 设置 PYSPARK_TIMEOUT + spark.hadoop.io.native.io.enabled=false |
| 修复 DLL 错误 | ✅ 删除 PATH 中 hadoop/bin,不加载 native IO |
| 超时问题 | ✅ 设置 PYSPARK_TIMEOUT=600,防止任务中断 |
| 小数据集 | ✅ 用 numSlices=1 控制分区 |
| 大数据集 | ✅ 用 repartition() 调优 |