教程原文:双Y轴截断图绘制教程
本期教程

本期教程,我们提供的原文的译文,若有需求请回复关键词:20240529
小杜的生信笔记 ,自2021年11月开始做的知识分享,主要内容是R语言绘图教程 、转录组上游分析 、转录组下游分析等内容。凡事在社群同学,可免费获得自2021年11月份至今全部教程,教程配备事例数据和相关代码,我们会持续更新中。
往期教程部分内容


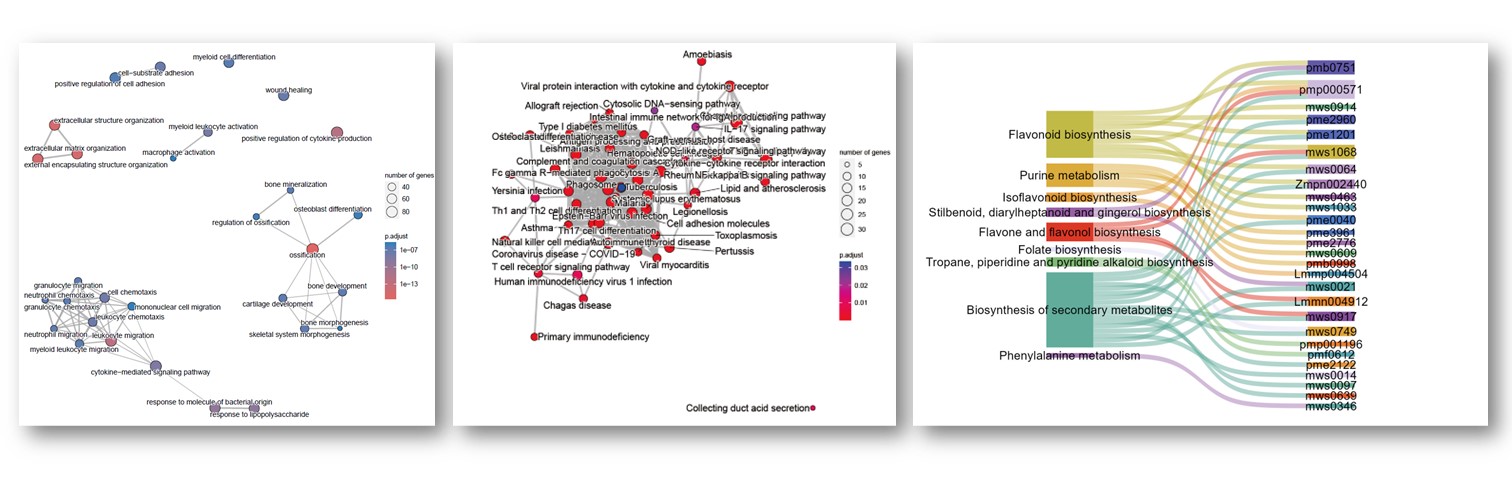
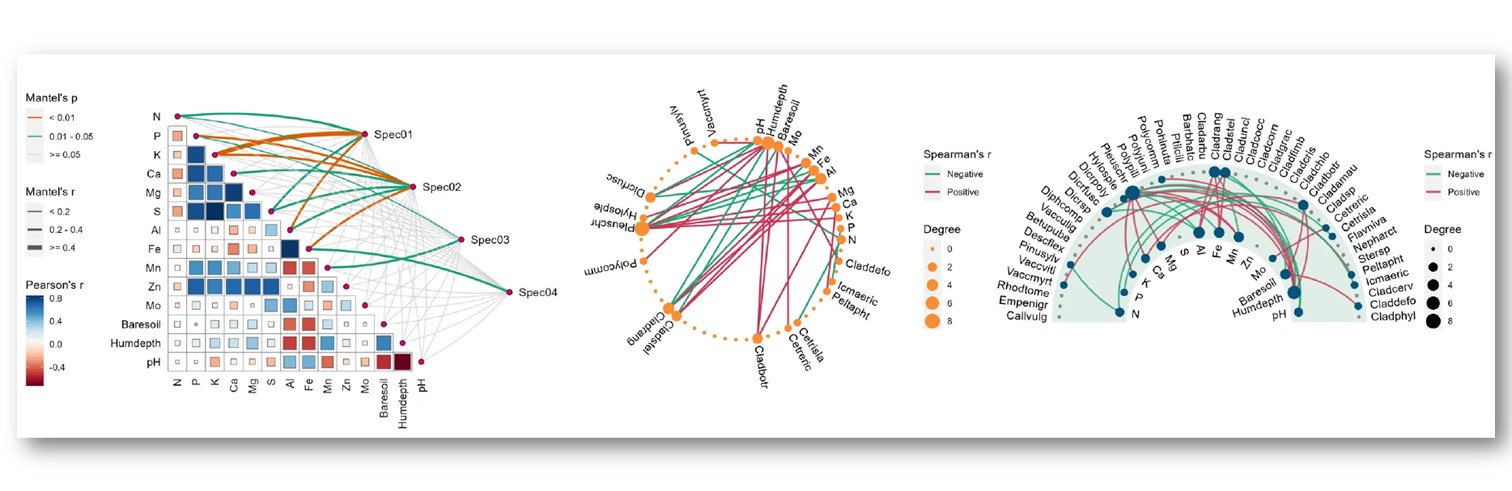

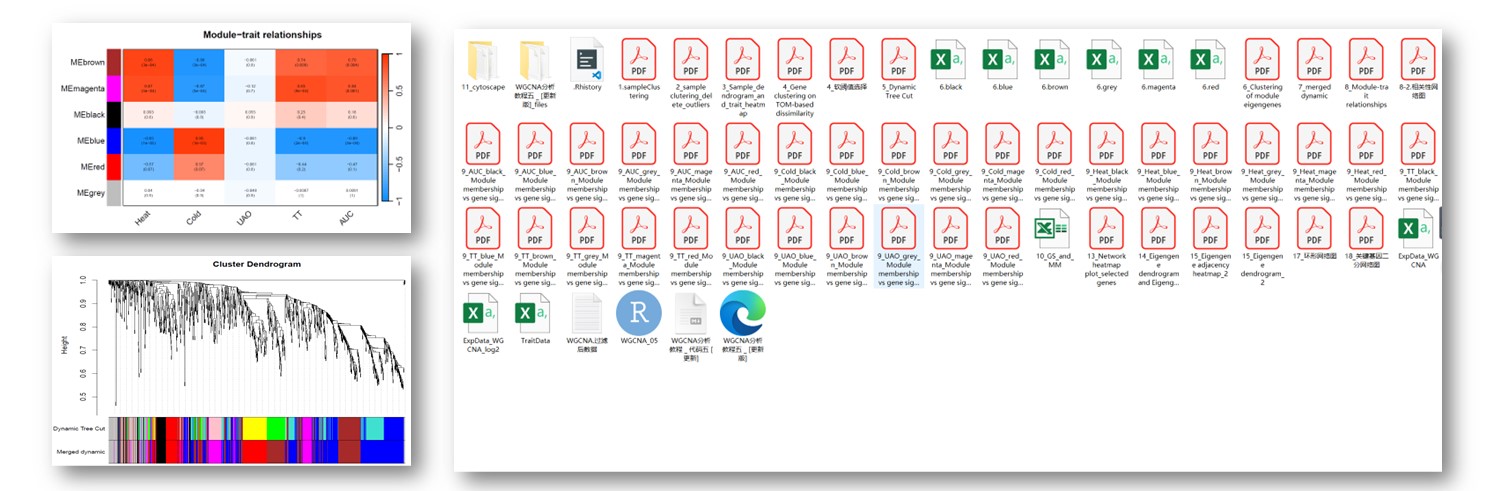

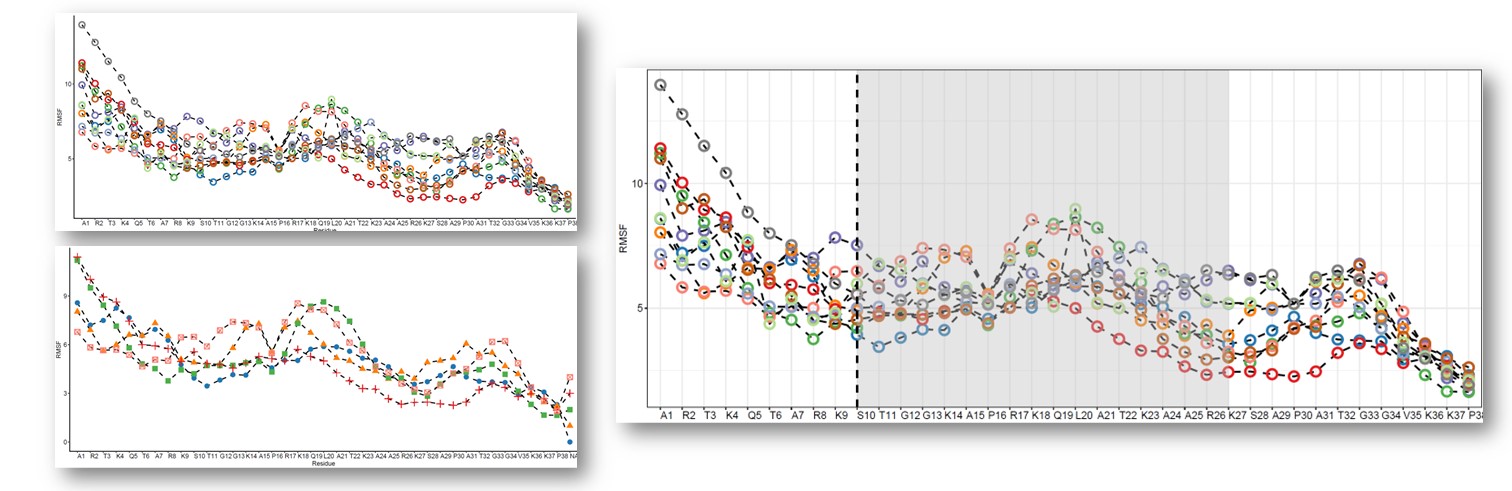
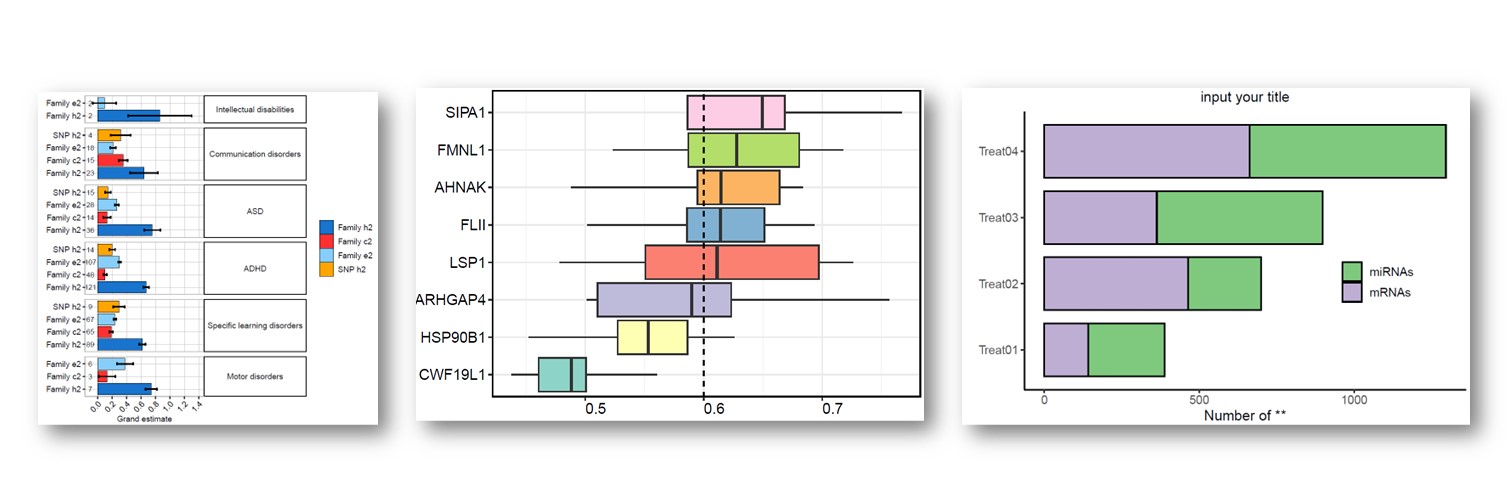
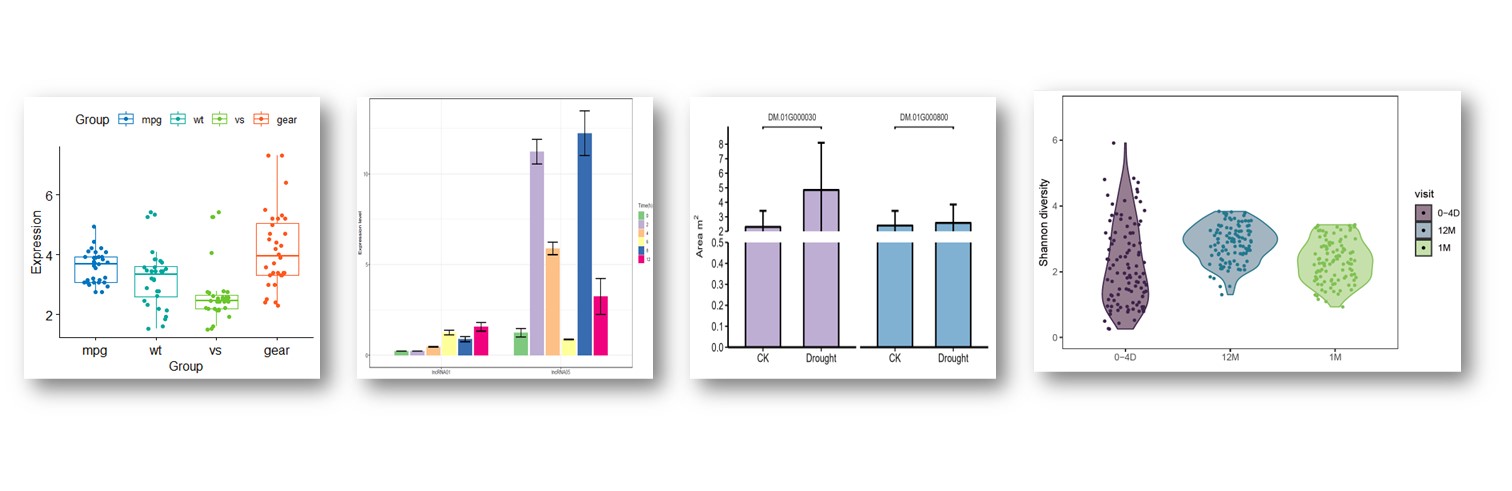
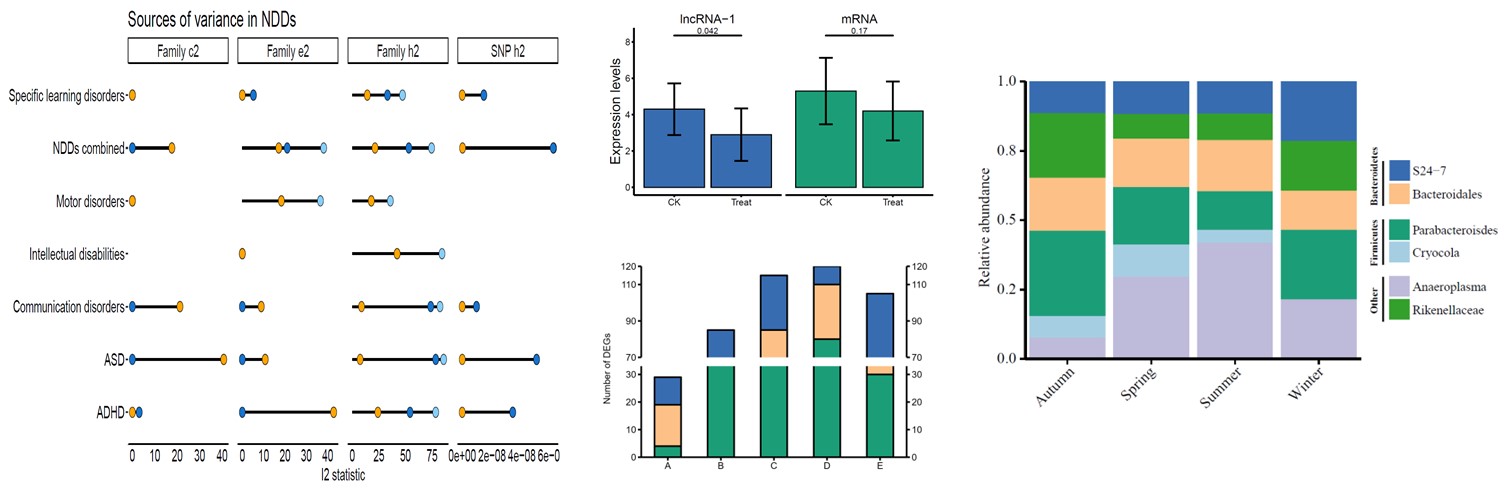
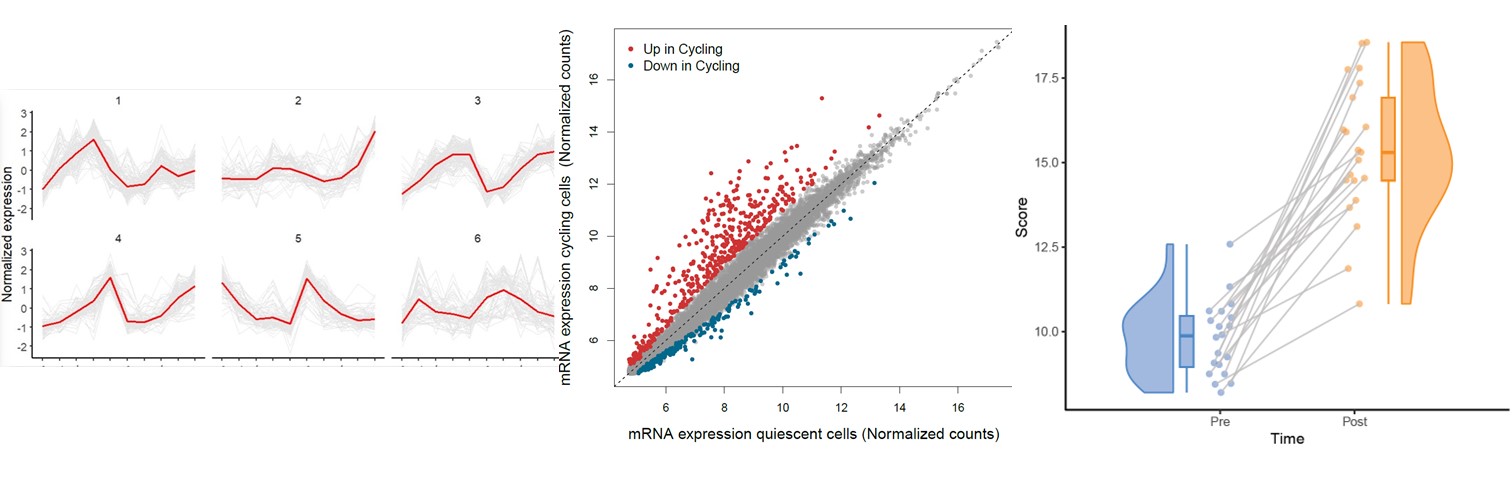

教程来源:
Yunyun Gao, Hao Luo, Yong-Xin Liu,et al, Benchmarking metagenomics tools for purging host contamination.绘图代码
R
library(ggplot2)
library(tidyverse)
library(ggbreak)
library(dunn.test)
library(car)
R
data <- read.table("data.txt", head = TRUE, sep = "\t")
R
# Select the relevant columns for data1
data1 <- data[, c("Software", "Taxa", "Rss")]
data2 <- data[, c("Software", "Taxa", "Time")]
data1$Software <- factor(data1$Software, levels = c( "BWA","Bowtie2","Kneaddata", "KMCP", "Kraken2", "Krakenuniq"))
R
ggplot(data1) +
geom_col(aes(x = Software, y = Rss, fill = Taxa), position = 'dodge', width = 0.8) +
scale_fill_manual(values = c('Rice' = '#fec79e', 'Human' = '#8ec4cb')) +
labs(x = "Software", y = "Memory usage / Gigabytes") +
theme_minimal() +
theme_test(base_size = 24) +
theme(
legend.position = 'none',
panel.border = element_rect(size = 2, fill = 'transparent'),
axis.text = element_text(color = 'black'),
axis.text.x = element_text(angle = 45, hjust = 1) # Rotate x-axis labels for better visibility
) +
geom_rect(aes(xmin = 0.5, xmax = 0.8, ymin = 53, ymax = 57), fill = '#8ec4cb', color = '#8ec4cb') +
geom_rect(aes(xmin = 1.4, xmax = 1.7, ymin = 53, ymax = 57), fill = '#fec79e', color = '#fec79e') +
annotate(geom = 'text', x = 1.9, y = 55, label = 'Rice', size = 7) +
annotate(geom = 'text', x = 1.1, y = 55, label = 'Human', size = 7) +
scale_y_continuous(
breaks = c(seq(0, 10, 10), seq(5, 10, 5), seq(20, 60, 10)),
limits = c(0, 60),
expand = c(0, 0),
sec.axis = sec_axis(~ .*5, name = 'Time consumption / minutes', breaks = c(seq(0, 50, 25), seq(50, 300, 50)))
) +
geom_point(data = data2, aes(x = factor(Software), y = Time * 0.2, color = Taxa, group = Taxa), shape=17, size = 5, na.rm = TRUE) +
scale_color_manual(values = c('#1e8b9b', '#ff8c3e')) +
scale_y_break(c(6,11), space = 0, scales = 1.8)Check Time of all taxa, Normality test
R
shapiro_test <- by(data$Time, data$Taxa, shapiro.test)
# Extract p-values from each group's test results
p_values <- sapply(shapiro_test, function(x) x$p.value)
# Check if each group conforms to normal distribution
normal_data <- p_values > 0.05
if (all(normal_data)) {
summary_stats <- aggregate(Time ~ Taxa, data = data, FUN = function(x) c(mean = mean(x), se = sd(x)/sqrt(length(x))))
print(summary_stats)
} else {
summary_stats <- aggregate(Time ~ Taxa, data = data, FUN = function(x) c(median = median(x), p25 = quantile(x, 0.25), p75 = quantile(x, 0.75)))
print(summary_stats)
}
#> Taxa Time.mean Time.se
#> 1 Human 99.474900 35.348141
#> 2 Rice 8.478217 2.883665
levene_test_result <- leveneTest(Time ~ Taxa, data = data)
p_value_levene <- levene_test_result$`Pr(>F)`[1]
if (all(normal_data)) {
if (p_value_levene > 0.05) {
t_test_result <- t.test(Time ~ Taxa, data = data, paired = TRUE)
print(t_test_result)
} else {
wilcox_result <- wilcox.test(Time ~ Taxa, data = data, paired = TRUE)
print(wilcox_result)
}
} else {
wilcox_result <- wilcox.test(Time ~ Taxa, data = data, paired = TRUE)
print(wilcox_result)
}
#>
#> Wilcoxon signed rank exact test
#>
#> data: Time by Taxa
#> V = 21, p-value = 0.03125
#> alternative hypothesis: true location shift is not equal to 0
R
shapiro_test <- by(data$Rss, data$Taxa, shapiro.test)
# Extract p-values from each group's test results
p_values <- sapply(shapiro_test, function(x) x$p.value)
# Check if each group conforms to normal distribution
normal_data <- p_values > 0.05
if (all(normal_data)) {
summary_stats <- aggregate(Rss ~ Taxa, data = data, FUN = function(x) c(mean = mean(x), se = sd(x)/sqrt(length(x))))
print(summary_stats)
} else {
summary_stats <- aggregate(Rss ~ Taxa, data = data, FUN = function(x) c(median = median(x), p25 = quantile(x, 0.25), p75 = quantile(x, 0.75)))
print(summary_stats)
}
#> Taxa Rss.median Rss.p25.25% Rss.p75.75%
#> 1 Human 5.6300000 4.8150000 18.2650000
#> 2 Rice 0.8040332 0.5614160 2.1725659
levene_test_result <- leveneTest(Rss ~ Taxa, data = data)
p_value_levene <- levene_test_result$`Pr(>F)`[1]
if (all(normal_data)) {
if (p_value_levene > 0.05) {
t_test_result <- t.test(Rss ~ Taxa, data = data, paired = TRUE)
print(t_test_result)
} else {
wilcox_result <- wilcox.test(Rss ~ Taxa, data = data, paired = TRUE)
print(wilcox_result)
}
} else {
wilcox_result <- wilcox.test(Rss ~ Taxa, data = data, paired = TRUE)
print(wilcox_result)
}
#>
#> Wilcoxon signed rank exact test
#>
#> data: Rss by Taxa
#> V = 21, p-value = 0.03125
#> alternative hypothesis: true location shift is not equal to 0教程原文:双Y轴截断图绘制教程
若我们的教程对你有所帮助,请
点赞+收藏+转发,这是对我们最大的支持。
往期部分文章
1. 最全WGCNA教程(替换数据即可出全部结果与图形)
2. 精美图形绘制教程
3. 转录组分析教程
4. 转录组下游分析
小杜的生信筆記 ,主要发表或收录生物信息学教程,以及基于R分析和可视化(包括数据分析,图形绘制等);分享感兴趣的文献和学习资料!!