基于sympy构造符号神经网络
本文的代码来自linjing-lab/optimtool,分类任务的代码在这里,回归任务的代码在这里,通用符号神经网络的代码在这里。
python
pip install optimtool>=2.8.0分类任务示例
python
import optimtool.unconstrain as ou
from optimtool.base import sp, np
w11, w21, w31 = sp.symbols('w11 w21 w31', real=True)
w12, w22, w32 = sp.symbols('w12 w22 w32', real=True)
b1, b2, b3 = sp.symbols('b1 b2 b3', real=True)
X = np.array([
[1.0, 1.0],
[1.5, 1.0],
[1.0, 1.5],
[4.0, 1.0],
[4.5, 1.0],
[4.0, 1.5],
[1.0, 4.0],
[1.5, 4.0],
[1.0, 4.5],
])
mean = X.mean(axis=0)
std = X.std(axis=0)
std[std == 0] = 1
X = (X - mean) / std
Y = np.zeros((X.shape[0], 3))
Y[:3, 0] = 1
Y[3:6, 1] = 1
Y[6:, 2] = 1
def symbolic_loss():
loss = 0.0
for i in range(len(Y)):
x1, x2 = X[i]
z1 = w11 * x1 + w12 * x2 + b1
z2 = w21 * x1 + w22 * x2 + b2
z3 = w31 * x1 + w32 * x2 + b3
exp_sum = sp.exp(z1) + sp.exp(z2) + sp.exp(z3)
p1 = sp.exp(z1) / exp_sum
p2 = sp.exp(z2) / exp_sum
p3 = sp.exp(z3) / exp_sum
loss_expr = - (Y[i, 0] * sp.log(p1) + Y[i, 1] * sp.log(p2) + Y[i, 2] * sp.log(p3))
loss += loss_expr
return loss / len(Y)
f_sym = symbolic_loss()
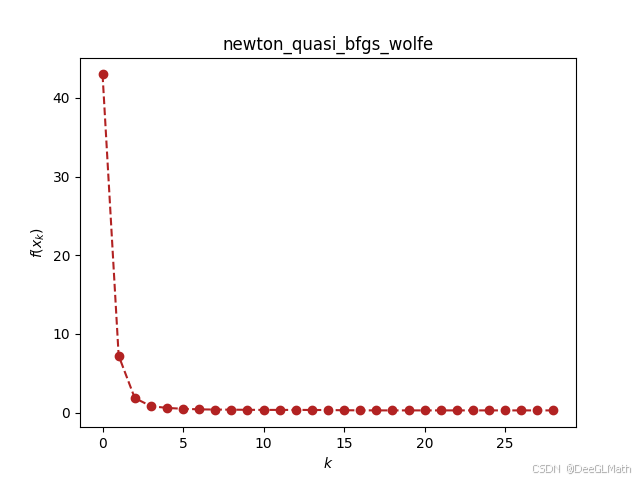
ou.gradient_descent.barzilar_borwein(f_sym, [w11, w21, w31, w12, w22, w32, b1, b2, b3], [0, 0, 0, 0, 0, 0, 0, 0, 0], verbose=True, epsilon=1e-2)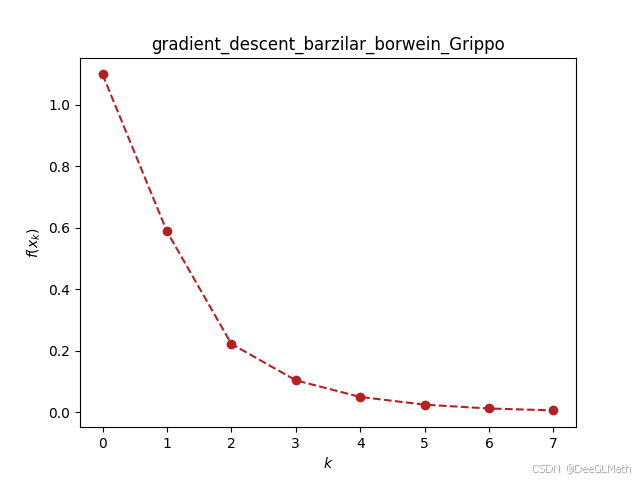
回归任务示例
python
import optimtool.unconstrain as ou
from optimtool.base import sp, np
w1, w2, w3, b = sp.symbols('w1 w2 w3 b', real=True)
np.random.seed(0)
n = 20 # samples
x1 = np.linspace(-3, 3, n)
x2 = 0.8 * x1 + np.random.randn(n) * 0.5
X = np.column_stack([x1, x2])
y_true = 2 * x1 - 3 * x2 + 1.5 * x1**2
noise = np.random.randn(n)
y = y_true + noise
def symbolic_loss():
loss = 0
for i in range(len(y)):
x1_, x2_ = X[i]
y_pred = w1 * x1_ + w2 * x2_ + w3 * x1_**2 + b
loss += (y_pred - y[i]) ** 2
return loss / len(y)
f_sym = symbolic_loss()
ou.newton_quasi.bfgs(f_sym, [w1, w2, w3, b], [0, 0, 0, 0], verbose=True)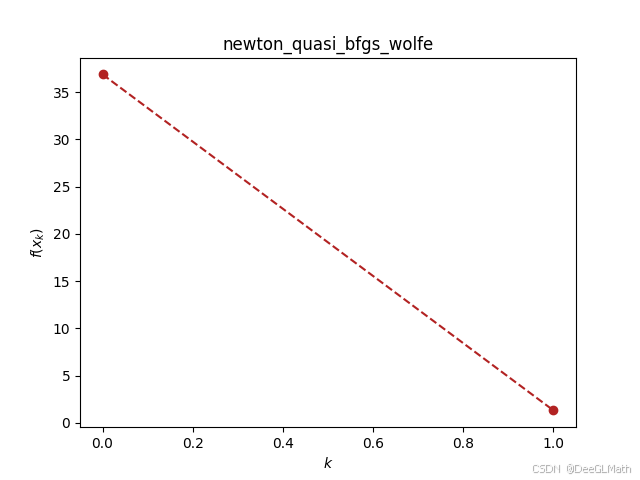
通用的神经网络
python
import optimtool.unconstrain as ou
from optimtool.base import sp, np
def gen_nn(X_data, y_data, hidden_dims=[], task='classification'):
n_samples, n_features = X_data.shape
if task == 'classification':
if len(y_data.shape) == 1:
n_outputs = len(np.unique(y_data))
y_one_hot = np.zeros((n_samples, n_outputs))
for i, label in enumerate(y_data):
y_one_hot[i, int(label)] = 1
else:
n_outputs = y_data.shape[1]
y_one_hot = y_data
y_processed = y_one_hot
elif task == 'regression':
if len(y_data.shape) == 1:
n_outputs = 1
y_processed = y_data.reshape(-1, 1)
else:
n_outputs = y_data.shape[1]
y_processed = y_data
else:
raise ValueError(f"Support classification or regression. not support {task}.")
params = []
layer_dims = [n_features] + hidden_dims + [n_outputs]
for i in range(len(layer_dims)-1):
prev_dim = layer_dims[i]
curr_dim = layer_dims[i+1]
for j in range(curr_dim):
for k in range(prev_dim):
params.append(sp.symbols(f'W{i}_{j}_{k}', real=True))
for j in range(curr_dim):
params.append(sp.symbols(f'b{i}_{j}', real=True))
def forward(X_vec):
idx = 0
x = X_vec.copy()
current_dim = n_features
for i, h_dim in enumerate(hidden_dims):
W_mat = []
for j in range(h_dim):
row = []
for k in range(current_dim):
row.append(params[idx])
idx += 1
W_mat.append(row)
b_vec = []
for j in range(h_dim):
b_vec.append(params[idx])
idx += 1
z = []
for j in range(h_dim):
sum_val = 0
for k in range(current_dim):
sum_val += W_mat[j][k] * x[k]
sum_val += b_vec[j]
z.append(sp.Max(0, sum_val)) # ReLU
x = z
current_dim = h_dim
output_dim = n_outputs
W_mat = []
for j in range(output_dim):
row = []
for k in range(current_dim):
row.append(params[idx])
idx += 1
W_mat.append(row)
b_vec = []
for j in range(output_dim):
b_vec.append(params[idx])
idx += 1
out = []
for j in range(output_dim):
sum_val = 0
for k in range(current_dim):
sum_val += W_mat[j][k] * x[k]
sum_val += b_vec[j]
out.append(sum_val)
return out
f_sym = 0
epsilon = 1e-10
if task == 'classification':
for s in range(n_samples):
logits = forward(X_data[s].tolist())
y_true = y_processed[s].tolist()
log_sum_exp = sp.log(sum(sp.exp(l) for l in logits) + epsilon)
probs = [sp.exp(l - log_sum_exp) for l in logits]
for c in range(n_outputs):
f_sym += -y_true[c] * sp.log(probs[c] + epsilon) # Softmax
f_sym /= n_samples
else: # regression
predictions = []
for s in range(n_samples):
pred = forward(X_data[s].tolist())
predictions.append(pred)
for s in range(n_samples):
for o in range(n_outputs):
diff = predictions[s][o] - y_processed[s, o]
f_sym += diff ** 2 # MSE
f_sym /= (n_samples * n_outputs)
return f_sym, params
# # classification
# X = np.array([
# [2.0, 1.0, 0.5, 0.1],
# [5.1, 3.1, 2.1, 1.05],
# [5.0, 3.0, 2.0, 1.0],
# [2.1, 0.9, 0.55, 0.15],
# [8.0, 5.0, 4.0, 2.0],
# [8.2, 5.2, 4.2, 2.1],
# ], dtype=np.float64)
# y = np.array([0, 1, 1, 0, 2, 2], dtype=np.int64)
# f_sym, params = gen_nn(X, y, hidden_dims=[2,])
# ou.gradient_descent.barzilar_borwein(f_sym, params, np.ones(len(params)).tolist(), verbose=True, epsilon=1e-2)
# regression
def gen_reg(n_samples=20, seed=0):
np.random.seed(seed)
X = np.linspace(-5, 5, n_samples).reshape(-1, 1)
noise = 0.2 * np.random.randn(n_samples, 3)
y1 = np.sin(X)
y2 = 0.5 * X
y3 = np.cos(X) + 0.2 * X
y = np.hstack([y1, y2, y3]) + noise
return X, y
X, y = gen_reg(n_samples=5)
f_sym, params = gen_nn(X, y, hidden_dims=[2,], task='regression')
ou.newton_quasi.bfgs(f_sym, params, np.ones(len(params)).tolist(), verbose=True, epsilon=1e-4)